Actin Cytoskeleton and Mechanotransduction
Mechanotransduction is a multi-step biological process by which cells sense, interpret, and respond to mechanical (i.e., physical) force through conversion to biochemical signals that elicit specific cellular responses. The responses are often mechanical in nature as they involve force generation to produce cellular protrusions and retractions which require remodeling of the actin cytoskeleton, consisting of monomeric (globular; G-) and helical polymeric (filamentous; F-) actin and actin binding proteins (ABPs)1-3. ABPs dynamically organize F-actin into many different structural forms such as lamellipodia, stress fibers, filopodia, podosomes, actin asters, vortices, and stars2-4. These different architectures serve specialized roles in the cell’s multiplex response to mechanical stimulation. A primary means by which F-actin transduces these signals is through its connections to focal adhesions and adherens junctions, which coordinate contact between the cell’s actin cytoskeleton and either the extracellular matrix or another cell, respectively5-7(Fig. 1). Understanding the actin cytoskeleton’s role in mechanotransduction goes beyond the basic biology underlying force-induced changes in actin-based cellular structures and functions. Diseases resulting from expression of mutant ABPs render cells unable to respond to mechanical forces physiologically3,8-13. In this newsletter, the role of the actin cytoskeleton in mechanotransduction is discussed.
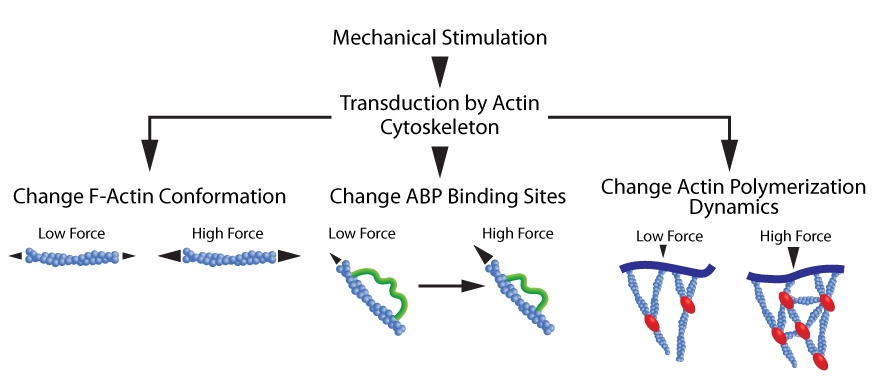
Figure 1. Actin cytoskeleton transduces mechanical forces. Mechanical loads induce a: 1. Conformational change in F-actin (left schematic); 2. Conformational change in ABPs that uncovers previously concealed binding sites (middle schematic); and 3. Alterations in ABP-mediated actin polymerization dynamics (right schematic).
The actin cytoskeleton functions as a mechanosensor for tension applied to cells3. The question is, how does the cytoskeleton respond to mechanical tension (Fig. 1)? Actin filaments in the various actin-based structures bear a mechanical load (per filament) that has been studied by a variety of microscopy techniques such as electron microscopy, Förster resonance energy transfer (FRET), atomic force microscopy, and optical traps, to name but a few3,14. These different actin structures are associated with specific mechanical loading that are optimized for the structure’s specialized cellular functions. Mechanical loading (increased tension) of filaments alters their conformation15 and how ABPs bind and affect filaments3. In the case of cofilin, a F-actin severing protein, changes in filament length affect its binding and function. Tensile forces that stretch a cell correspondingly increase the length of filaments parallel to the direction of the stretch. Under these conditions, the binding affinity of cofilin is reduced and that of myosin II is increased16-18. This mechanical-induced change in F-actin length and binding partners results in stabilized F-actin which can more easily form stress fibers, an essential part of a cell’s mechanotransduction processes7,16-19 (Fig. 1). Tension-induced changes in actin structural dynamics also affects the binding of actin-nucleating proteins such as Arp2/320. Mechanically-induced changes in actin-based structures can also affect gene expression in at least some cell types. As more stress fibers form during mechanical stimulation, the transcriptional coactivator YAP translocates to the nucleus where it is activated. YAP is integral in Hippo signaling and mediates increased expression of genes involved in cell proliferation and differentiation. Thus, the response of the actin cytoskeleton to extracellular mechanical forces can result in processes that have both physiological and pathophysiological relevance7,21,22.
Similar to F-actin, ABPs and other actin-associated proteins directly respond to mechanical stresses (Fig. 1). Common responses include conformational changes which expose previously concealed protein binding sites. This is the case for talin, a focal adhesion-associated protein, the adherens junction protein α-catenin, and ABPs such as vinculin, filamins, myosin, α-actinin 4, and actin filament-associated protein23-29.
Finally, actin polymerization and network assembly are modulated by mechanical forces (Fig. 1). Through force-induced changes in ABPs (see above), polymerization dynamics are altered3. By itself, mechanical force itself can also oppose polymerization by acting as a physical obstacle. Such changes in polymerization dynamics can alter the density and organization of filaments3.
Summary
Actin is the quintessential cytoskeletal protein and perhaps is the protein most associated with a cell’s response to varied external stimuli that results in changes to cellular shape, motility, intracellular trafficking, and force generation. Despite actin’s obvious importance in physiological and pathophysiological processes and decades of focused research, challenges remain in understanding how so many different higher order actin structures exist in a cell and what their corresponding functions are in mechanotransduction. Answering these questions is technically challenging as it requires high resolution microscopy combined with applying and measuring the mechanical load on filaments. To help scientists unravel the roles of F-actin in mechanotransduction (and other cellular processes), Cytoskeleton, Inc. offers purified labeled and unlabeled actin proteins, purified ABPs, functional actin-based assay kits, and F-actin live cell imaging probes.
References
- Alonso J.L. and Goldmann W.H. 2016. Cellular mechanotransduction. AIMS Biophysics. 3, 50-62.
- Wang N. 2017. Review of cellular mechanotransduction. J. Phys. D. Appl. Phys. 50(23). pii: 233002.
- Harris A.R. et al. 2018. Mechanotransduction by the actin cytoskeleton: Converting mechanical stimuli into biochemicals signals. Annu. Rev. Biophys. 47, 617-631.
- Fritzsche M. et al. 2017. Self-organizing actin patterns shape membrane architecture but not cell mechanics. Nat. Commun. 8, 14347.
- Perez-Moreno M. et al. 2003. Sticky business: orchestrating cellular signals at adherens junctions. Cell. 112, 535-548.
- Geiger B. et al. 2009. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21-33.
- Ohashi K. et al. 2017. Roles of the cytoskeleton, cell adhesion and rho signaling in mechanosensing and mechanotransduction. J. Biochem. 161, 245-254.
- Jaalouk D.E. and Lammerding J. 2009. Mechanotransduction gone awry. Nat. Rev. Mol. Cell Biol. 10, 63-73.
- Weins A. et al. 2007. Disease-associated mutant α-actinin-4 reveals a mechanism for regulating its F-actin-binding affinity. Proc. Natl. Acad. Sci. USA. 104, 16080-16085.
- Lee S.H. et al. 2008. Crystal structure of the actin-binding domain of α-actinin-4 Lys255Glu mutant implicated in focal segmental glomerulosclerosis. J. Mol. Biol. 376, 317-324.
- Clark A.R. et al. 2009. Skeletal dysplasias due to filamin A mutations result from a gain-of-function mechanism distinct from allelic neurological disorders. Hum. Mol. Genet. 18, 4791-4800.
- Henderson D.M. et al. 2009. Patients with ACTN4 mutations demonstrate distinctive features of glomerular injury. J. Am. Soc. Nephrol. 20, 961-968.
- Henderson D.M. et al. 2010. Disease-causing missense mutations in actin binding domain 1 of dystrophin induce thermodynamic instability and protein aggregation. Proc. Natl. Acad. Sci. USA. 107, 9632-9637.
- Wang Y. and Kanchanawong P. 2016. Three-dimensional super resolution microscopy of F-actin filaments by interferometric photoactivated localization microscopy (iPALM). JoVE. 118, e54774.
- Shimozawa T. and Ishiwata S. 2009. Mechanical distortion of single actin filaments induced by external force: detection by fluorescence imaging. Biophys. J. 96, 1036-1044.
- Uyeda T.Q. et al. 2011. Stretching actin filaments within cells enhances their affinity for the myosin II motor domain. PLoS One. 6, e26200.
- Hayakawa K. et al. 2011. Actin filaments function as a tension sensor by tension-dependent binding of cofilin to the filament. J. Cell Biol. 195, 721-727.
- Hayakawa K. et al. 2014. Single-molecule imaging and kinetic analysis of cooperative cofilin-actin filament interactions. Proc. Natl. Acad. Sci. USA. 111, 9810-9815.
- McGough A. et al. 1997. Cofilin changes the twist of F-actin: implications for actin filament dynamics and cellular function. J. Cell Biol. 138, 771-781.
- Risca V.I. et al. 2012. Actin filament curvature biases branching direction. Proc. Natl. Acad. Sci. USA. 109, 2913-2918.
- Dupont S. et al. 2011. Role of YAP/TAZ in mechanotransduction. Nature. 474, 179-183.
- Halder G. et al. 2012. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591-600.
- Han B. et al. 2004. Conversion of mechanical force into biochemical signaling. J. Biol. Chem. 279, 54793-54801.
- del Rio A. et al. 2009. Stretching single talin rod molecules activates vinculin binding. Science. 323, 638-641.
- Ehrlicher A.J. et al. 2011. Mechanical strain in actin networks regulates FilGAP and integrin binding to filamin A. Nature. 478, 260.
- Luo T. et al. 2013. Molecular mechanisms of cellular mechanosensing. Nat. Mater. 12, 1064-1071.
- Buckley C.D. et al. 2014. The minimal cadherin-catenin complex binds to actin filaments under force. Science. 346, 1254211.
- Huelsmann S. et al. 2016. Evidence for the mechanosensory function of filamin in tissue development. Sci. Rep. 6, 32798.
- Schiffhauer E.S. et al. 2016. Mechanoaccumulative elements of the mammalian actin cytoskeleton. Curr. Biol. 26, 1473-1479.
Related Products
Actin Products
Actin protein (>99% pure): bovine cardiac muscle (Cat. # AD99)
Actin protein (>99% pure): chicken gizzard muscle (Cat. # AS99)
Actin protein (pre-formed filaments): rabbit skeletal muscle (Cat. # AKF99)
Actin protein (>95% pure): rabbit skeletal muscle (Cat. # AKL95)
Actin protein (>99% pure): rabbit skeletal muscle (Cat. # AKL99)
Actin protein (>99% pure): human platelet (Cat. # APHL99)
Actin protein (rhodamine): human platelet (Cat. # APHR)
Actin protein (rhodamine): rabbit skeletal muscle (Cat. # AR05)
Spirochrome SiR-Actin Kit (Cat. # CY-SC001)
Spirochrome SiR700-Actin Kit (Cat. # CY-SC013)
Acti-stain 488 Phalloidin (Cat. # PHDG1)
Acti-stain 555 Phalloidin (Cat. # PHDH1)
Acti-stain 670 Phalloidin (Cat. # PHDN1)
Rhodamine Phalloidin (Cat. # PHDR1)
Actin Biochem Kits
Actin Binding Protein Spin-Down Assay Biochem Kit: rabbit skeletal muscle actin (Cat. # BK001)
Actin Binding Protein Spin-Down Assay Biochem Kit: human platelet actin (Cat. # BK013)
Actin Polymerization Biochem Kit (fluorescence format): rabbit skeletal muscle actin (Cat. # BK003)

