Actin protein (pyrene labeled): rabbit skeletal muscle
Product Uses Include
- Studying actin polymerization in vitro.
- Studying the effects of actin binding proteins and drugs on actin polymerization in vitro.
Material
Purified rabbit muscle actin (Cat. # AKL99) has been modified to contain covalently linked pyrene at the cysteine 374 residue. An N-(1-pyrene) iodoacetamide is used to label the actin protein. Pyrene labeling stoichiometry has been determined to be 0.6 dyes per actin monomer. Pyrene labeled rabbit muscle actin has an approximate molecular weight of 43 kDa, and is supplied as a white lyophilized powder. The lyophilized protein is st able for 6 months when stored desiccated to <10% humidity at 4°C. The protein should be reconstituted to 20 mg/ml with distilled water; it will then be in the following buffer: 5 mM Tris-HCl pH 8.0, 0.2 mM CaCl2, 0.2 mM ATP, 5% sucrose, and 1% dextran.
Purity
Protein purity is determined by scanning densitometry of Coomassie Blue stained protein on a 4-20% gradient polyacrylamide gel. Pyrene muscle actin is >99% pure (see Figure 1).
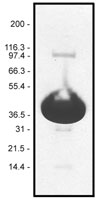
Figure 1. Pyrene Muscle Actin Protein Purity Determination. A 100 µg sample of pyrene muscle actin (molecular weight approx. 43 kDa) was separated by electrophoresis in a 4-20% SDS-PAGE system, and stained with Coomassie Blue. Protein quantitation was determined with the Precision Red Protein Assay Reagent (Cat. # ADV02).
Biological Activity
The fluorescent signal of monomer pyrene actin is enhanced during its polymerization into filaments, making it an ideal tool for monitoring actin filament formation. Stringent quality control ensures that AP05 pyrene F-actin has a 7-12 fold fluorescent enhancement over non-polymerized pyrene G-actin (See Fig 2).
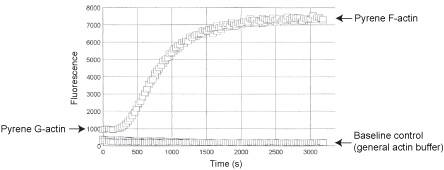
Figure 2. Fluorescence enhancement during pyrene actin polymerization. Pyrene muscle actin was polymerized in duplicate wells of a 96-well plate by the addition of Actin Polymerization Buffer (Cat. # BSA02). The fluorescent signal was scanned every 30 s for 1 h. Polymerized pyrene F-actin shows a 10 fold fluorescent enhancement over non-polymerized pyrene G-actin and buffer control.
For product Datasheets and MSDSs please click on the PDF links below. For additional information, click on the FAQs tab above or contact our Technical Support department at tservice@cytoskeleton.com
Question 1: When performing the polymerization assay using pyrene-labeled actin (Cat. # AP05), the fluorescence signal of the globular actin in G-buffer increased over time, so that the 0 value was already rather high.
Answer 1: After preparing actin at a concentration of 0.4 mg/ml for polymerization (from either fresh stocks or frozen aliquots), it is important to incubate the actin on ice for 1 hour to depolymerize any actin polymers that might have formed during storage. For an even cleaner sample of G-actin, the samples can be centrifuged (100,000 x g) for 60 min to absolutely insure the removal of residual actin oligomers and nucleating centers. After centrifugation, use the top 80% of the supernatant as your G-actin stock. This will eliminate any changes in fluorescence signal associated with G-actin.
Question 2: During my polymerization reaction, the growth phase and steady-state levels of F-actin produced are lower than what is shown in the manual. Why?
Answer 2: One of the most common causes of reduced fluorescence intensity signal is non-optimal machine settings. Please check that the machine’s settings are similar to these:
Measurement type: Kinetic 120 cycles, 60 sec interval time
Fluorescence wavelengths: Ex. 350 or 360 +/- 20 nm
Em. 407 or 410 +/- 10 nm or 420 +/- 20 nm
Gain: 100 (on a scale of 0-120, where 120 is the highest)
Reads per well: 1
Fluorescence reading from: Top
Also, when samples are not being measured in the fluorimeter, the excitation beam should be shuttered to prevent the beam from bleaching the pyrene fluorescence. When measuring the fluorescence, the shutter should be opened for no longer than 7 sec every 30 sec for 1 h.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com.