SOS1 Protein (ExD Exchange Domain, aa564-1049, 6xHis tag)
* Limited stock available. If stock is not available, Cytoskeleton will produce a new batch upon request. Minimum order will apply. Inquire for more information.
Product Uses
- Study inhibitors of SOS1 GTP exchange activity
- Identification of SOS1 binding proteins
- Study of SOS1 GEF activity with different GTPases
Materials
The human SOS1 exchange domain (564-1049) protein (SOS1-ExD) has been produced in a bacterial expression system. The recombinant protein contains a 6 histidine tag (His-tag) at its amino terminus. The approximate molecular weight of the SOS1-ExD protein is 61 kDa. The protein is supplied as a lyophilized white powder.
Storage
The protein should be reconstituted to 50 µM (3.03 mg/ml) by the addition of 33 μl of distilled water. The protein will be in the following buffer: 17 mM Tris pH 7.5, 17 mM NaCl, 0.2 mM MgCl2, 2.3% sucrose, and 0.3% dextran. In order to maintain high biological activity of the protein, it is strongly recommended that the protein solution be aliquoted into "experiment sized" aliquots and snap frozen in liquid nitrogen. The protein can be stored at -70°C for 6 months. Avoid repeated freeze-thaw cycles. The lyophilized protein is stable at 4°C for 1 year when stored desiccated (<10% humidity).
Purity
Protein purity is determined by scanning densitometry of Coomassie blue stained protein on a 4-20% polyacrylamide gradient gel. The SOS1-ExD protein was determined to be >90% pure. (see Figure 1).
Figure 1. His-hSOS1 (564-1049) Protein Purity Determination
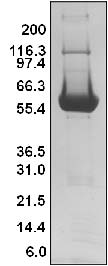
Legend: A 20 μg sample of recombinant SOS1-ExD protein (molecular weight approx. 61 kDa) was separated by electrophoresis in a 4-20% SDS-PAGE system. The protein was stained with Coomassie blue. Protein quantification was determined using the Precision Red Protein Assay Reagent (Cat.# ADV02). Mark12 molecular weight markers are from Invitrogen.
Biological Activity Assay
Human SOS1 is an exchange factor for the small G-protein Ras.The biological activity of SOS1 can be determined from its ability to catalyze the exchange of GDP for GTP on Ras. A standard biological assay for monitoring the catalytic activity of hSOS1 is an exchange assay utilizing the 2X Exchange Buffer from the RhoGEF exchange assay kit (Cat.# BK100).
Reagents
1. Recombinant Ras protein
2. Recombinant SOS1-ExD protein (Cat.# GE02)
3. 2x Exchange Buffer (40 mM Tris pH 7.5, 100 mM NaCl, 20 mM MgCl2 , 0.1 mg/ml BSA, 1.5 μM mant-GTP)
4. Dilution Buffer (20 mM Tris pH 7.5, 50 mM NaCl, 10 mM MgCl2, 0.1 mg/ml BSA)
Equipment
1. Fluorescence spectrometer (λex=360nm, λem=440nm)
2. Corning 96-well half area plates (Cat # 3686) or other plate with low protein binding surface.
Method
1. Dilute SOS1-ExD protein (Cat# GE02) to 6 μM (0.36 mg/ml) with Dilution Buffer.
2. Dilute Ras to 50 μM (1.1 mg/ml) with Dilution Buffer.
3. Dissolve lyophilized 2x Exchange Buffer in 5 ml Milli-Q water and keep at room temperature.
4. Set up the plate reader for kinetic fluorescence measurements (Excitation wavelength at 360 nm and emission wavelength at 440 nm) with readings every 30 seconds for 30 minutes.
5. Add the following components together and mix well by gentle pipeting:
Exchange reaction mix / 96 well black plate
2x Exchange Buffer / 50 μl
dH2O / 36 μl
50 μM Ras / 4 μl
6. Pipette 10 μl of 6 μM SOS1-ExD protein or Dilution Buffer in their respective wells and immediately pipette up and down twice and begin reading the fluorescence.
7. Once the readings are complete and the plate reader file has been saved, the exchange rate can be calculated by reducing the data to Vmax with the software that accompanies the plate reader.
Figure 2. SOS1-ExD protein mediated mant-GTP exchange on Ras.
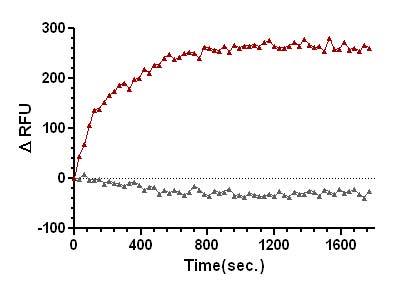
Legend: Ras protein was added to the wells of a 96-well half area plate containing diluted Exchange Buffer and mixed well. To initiate the exchange reaction, SOS1-ExD protein (red triangles), or Dilution Buffer (grey triangles), was added to the wells, mixed, and fluorescence measurements were obtained using a Tecan SpectraFluor Plus Spectrophotometer. The average fluorescence data from duplicate assays were normalized to the fluorescence at the zero time point. The resulting data were plotted as the change in relative fluorescence units (ΔRFU) over time using GraphPad Prism software.
For product Datasheets and MSDSs please click on the PDF links below.
Q1. What formats are available to study nucleotide exchange on small G-proteins?
A1. Several types of assay exist, these range from radio-labeled nucleotide filter based assays, to fluorescence enhancement using fluorescent nucleotide conjugates such as mant-GTP or -GDP, or Bodipy-FL-GTP or-GDP. These reporters enhance their fluorescence when binding to the small G-protein and hence the fluorescent signal is proportional to the amount of reporter-occupied nucleotide binding sites.
Q2. Which is best to measure nucleotide association or dissociation?
A2. Either can be used to indicate general nucleotide exchange, and the preference lies in whether you are looking for exchange for activation purposes, for example in the case of tumor physiology Ras GEFs are up-regulated which increases the GTP bound state of the small G-protein, thus in drug development efforts its important to know the rate of GDP dissociation and replacement with GTP, which is assumed to correlate with activation in cells.
Q3. What should be titrated to achieve a useful GTP exchange assay?
A3. The two factors that affect the signal to noise of a GEF assay are the proteins themselves, usually GEFs are tritrated from 50 nM to 5 µM, and small G-proteins from 0.5 to 5 µM.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com
