Ubiquitination Control Beads
As part of the Signal-Seeker™ product line, ubiquitination control beads (CUB02B-beads) have been developed as highly specific negative control beads for Cytoskeleton's ubiquitination affinity bead (UBA01B-Beads) products. These control beads are far superior to beads alone, because it has mutated versions of the UBDs covalently attached to the control bead matrix. This control is particularly important for ubiquitin IPs as the unmodified version of target proteins often bind non-specifically to all available ubiquitin enrichment affinity matrices. A comprehensive Signal-Seeker™ Ubiquitin Detection Kit (BK161), which includes ubiquitin control beads (CUB02B-beads), is available and is recommended for first time users.
Note: The product #s have been updated from UBA01 and CUB02 to UBA01B and CUB02B beads due to a reformulation of the bead preservation buffer. Reformulation results in an enhanced bead performance post-lyophilization. The affinity protein formulation on the beads has not changed.
Validation Data: Ubiquitination Control Beads White Paper
Each lot of affinity-bead is quality controlled to provide high batch to batch consistency, see COA documents.
Validated Applications
Application 1: Utilization of IgG control beads
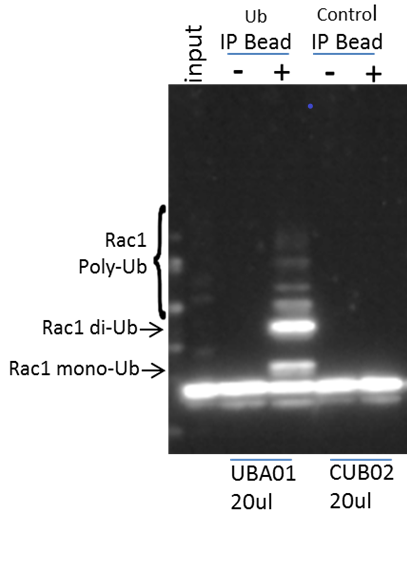
Figure 1: Swiss 3T3 cells were pre-treated with MG-132, then either untreated or treated with CNF1 (CN04) for 3 hours prior to lysis with BlastR buffer. The BK161 kit was utilized to perform the IP on 300 μg of lysate per condition. Input: 3 μg input untreated lysate, Ubiquitin Affinity beads (UBA01) plus untreated lysate, UBA01 plus CN04 treated lysate, ubiquitin control beads (CUB02) plus untreated lysate, and CUB02 plus CN04 treated lysate. Samples were analyzed for Rac1 ubiquitination using an anti-Rac1 antibody.
Amount:
Each package contains enough ubiquitin control beads for 10 reactions.
For more information contact: signalseeker@cytoskeleton.com
Associated Products:
Signal-Seeker™ Ubiquitination Detection Kit (Cat. # BK161)
Signal-Seeker™ Ubiquitin Affinity Beads (Cat.# UBA01B-beads)
Signal-Seeker™: BlastR™ Rapid Lysate Prep Kit (Cat. # BLR01)
Signal-Seeker™: PTMtrue™ Ubiquitin Antibody (Cat.# AUB01)
Visit our Signal-Seeker™ Tech Tips and FAQs page for technical tips and frequently asked questions regarding this and other Signal-Seeker™ products click here
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com