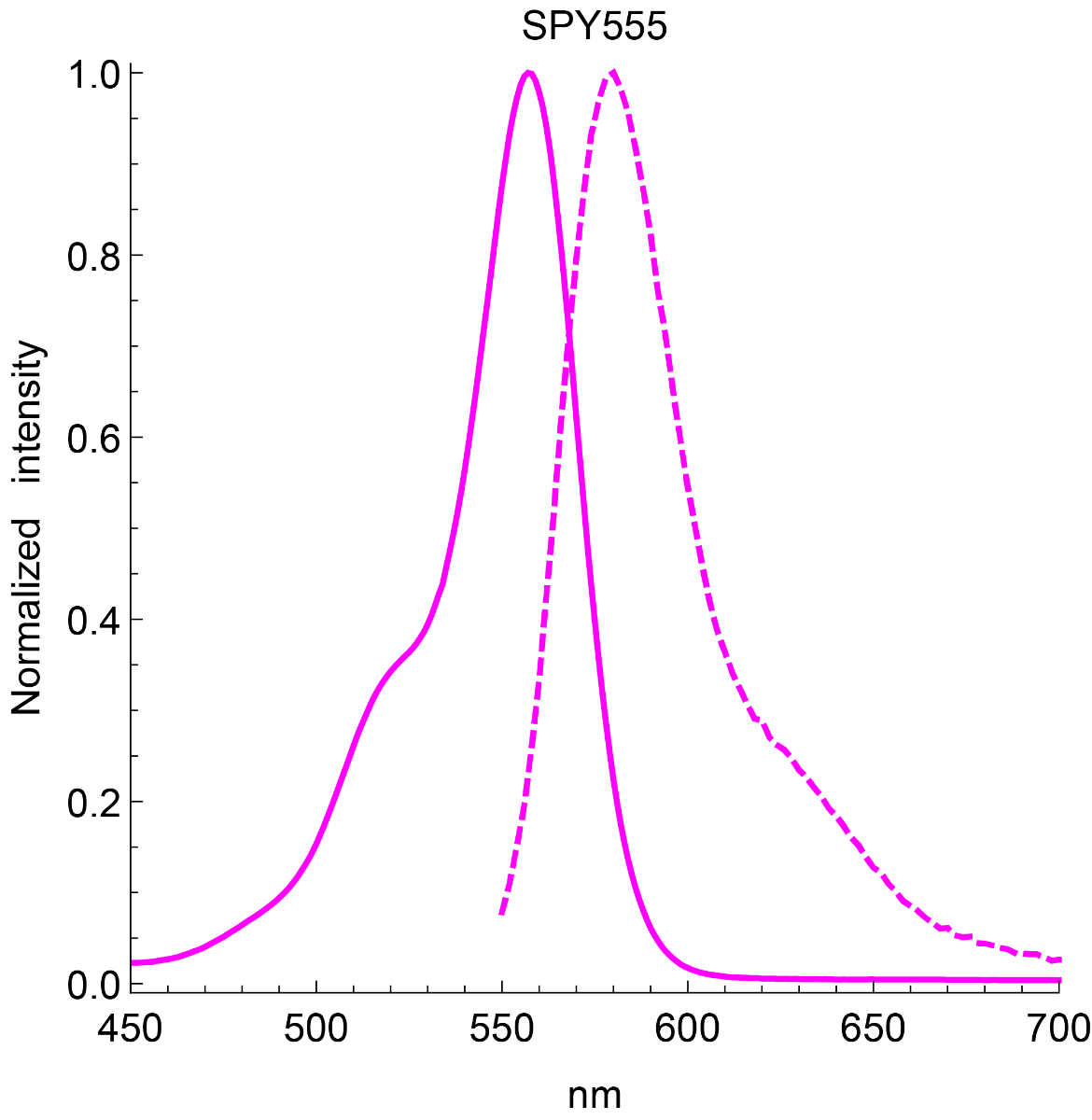
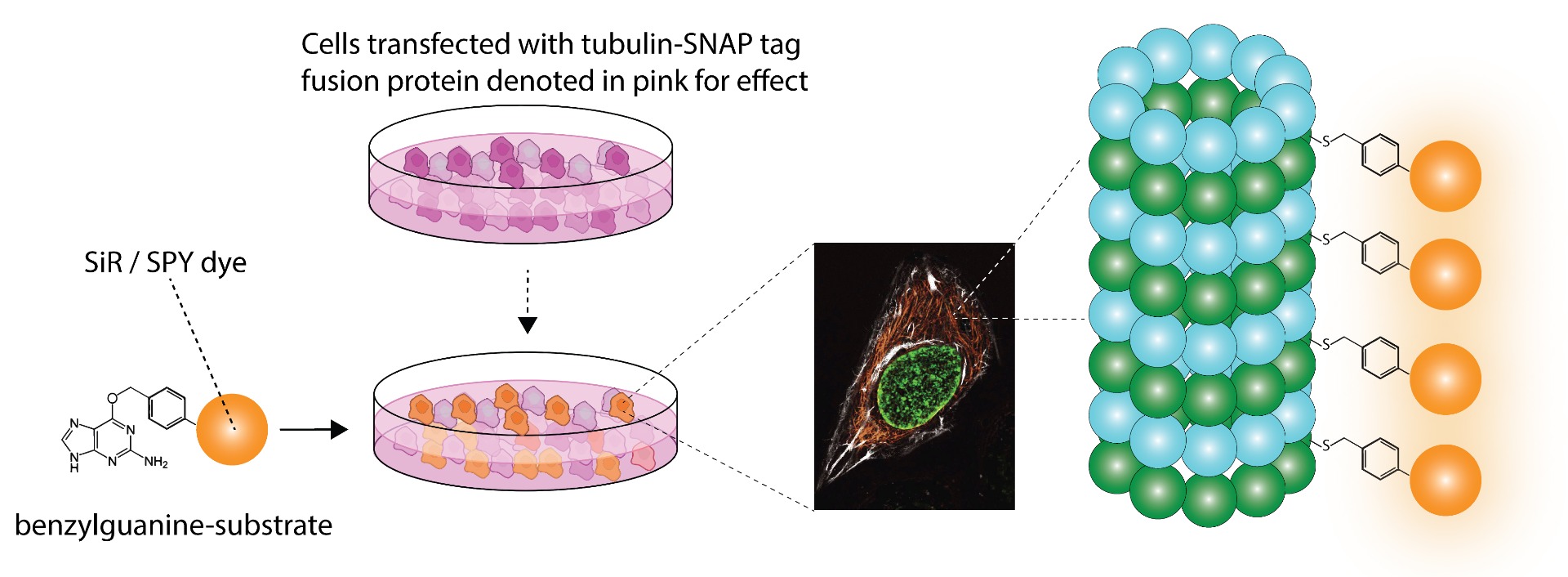
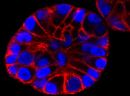
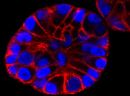
SPY555-BG is the benzylguanine derivative of our SPY555 fluorophore. It emits light in the orange part of the UV-ViS spectrum, it is fluorogenic, highly cell permeable and well suited for STED and SIM superresolution imaging. SPY555-BG can be imaged with a standard Cy5 filterset. It can be used for widefield, confocal, SIM or STED imaging in living or fixed cells and tissue. It has the same structure as MaP555-SNAP the name used in its original publication in Nature Chemistry1). SPY555-BG is an alternative to SNAP-Cell® TMR-Star, it has faster labeling kinetics.
Benzylguanine (BG) is the substrate of the self labeling tag SNAP-tag™*. Upon reaction with a BG derivative, SNAP-tag™* forms a covalent bond with the substrate and releases guanine. It allows to permanently attach a fluorescent label to any protein of interest (POI) expressed as SNAP-tag™* fusion.
*SNAP-tag™ and SNAP-Cell® are registered trademarks of New England Biolabs, Inc.
For product Datasheets and MSDSs please click on the PDF links below.
Spirochrome Technical Tips and Ex/Em spectra in graphical form (PDF)
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com
Q1. What is STED microscopy and how does it work?
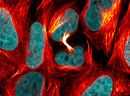
A1. STED microscopy stands for Stimulated Emission Depletion microscopy. It is one type of super resolution microscopy which allows the capture of images with a higher resolution than conventional light microscopy which is constrained by diffraction of light. STED uses 2 laser pulses, one is the excitation pulse which excites the fluorophore, causing it to fluoresce. The second pulse, referred to as the STED pulse, de-excites the fluorophore via stimulated emission in an area surrounding a central focal spot that is not de-excited and thus continues to fluoresce. This is accomplished by focusing the STED pulse into a ring shape, a so-called donut, where the center focal spot is devoid of the STED laser pulse, conferring high resolution to the fluorescent area (Fig. 1; see Ref. 1 for more details on STED microscopy).
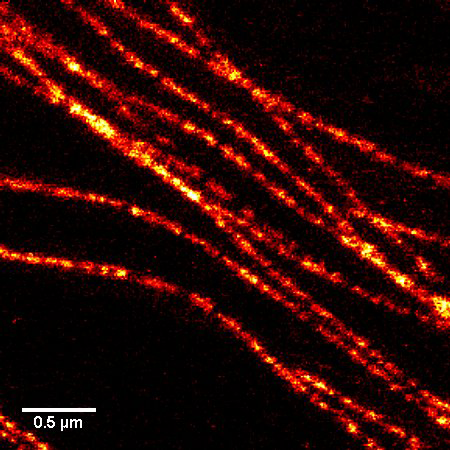
Figure 1. STED microscopic image of microtubules labeled with SiR-tubulin in human primary dermal fibroblasts.
Q2. Why is the SPY-BG and SiR-BG probe good for STED microscopy?
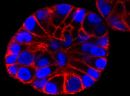
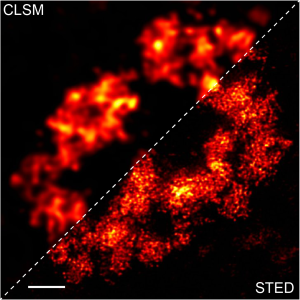
A2. STED microscopy offers the ability to study cellular details on a nanometermolar scale in vivo. To take advantage of this super resolution microscopy, one must be able to select with high specificity the area to be examined using fluorescent probes. In addition, the fluorescent probes must be bright, photostable, exhibit no or little phototoxicity, be excited and emit in the far red spectrum. In addition, if the probe is to be used for live cell imaging (thus avoiding fixation artifacts that occur when cells are fixed), high cell permeability is necessary. The SPY-BG and SiR-BG probes fulfill all of these requirements. As an example, the combination of STED and SiR probes allows for unparalleled fluorescent visualization of subcellular actin and tubulin/microtubule structures and their physical characterization in living cells, (see Fig. 2 and Ref. 2).
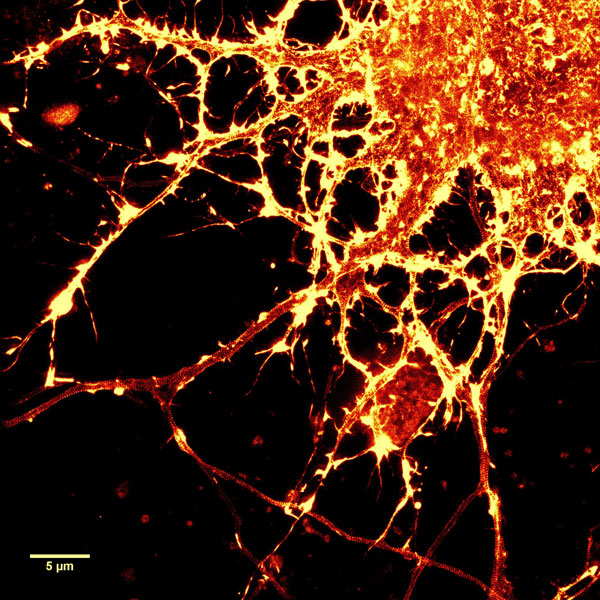
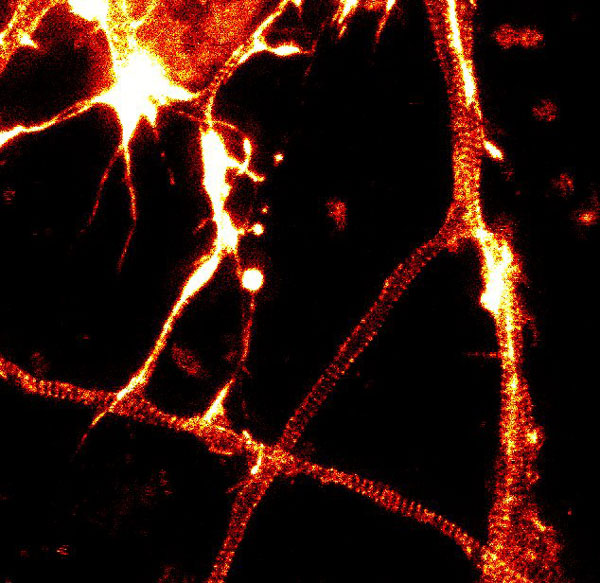
Figure 2. STED images of cultured rat hippocampal neurons stained with SiR-actin. Bottom image is a close-up view of part of the top image to clearly visualize actin rings (stripes) with 180 nm periodicity. Courtesy Of Elisa D'Este, MPI Biophysical Chemistry, Göttingen.
Q3: Are the SPY probes stable at room temperature?
A3: Yes, the probes are stable at room temperature for a few days. However, it strongly depends on the probe and the solvent. Thus, it is recommended to store all of the probes or solutions at –20°C.
References
1. Hell S.W. and Wichmann J. 1994. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780-782.
2. D’Este E. et al. 2015. STED nanoscopy reveals the ubiquity of subcortical cytoskeleton periodicity in living neurons. Cell Rep. 10, 1246-1251.
3. Lukinavicius G. et al. 2013. A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins. Nat. Chem. 5, 132-139.
4. Lukinavicius G. et al. 2014. Fluorogenic probes for live-cell imaging of the cytoskeleton.Nature Methods. 11, 731-733.