December 2019 Newsletter: Phenotypic Profiling: Actin-focused Cancer Therapeutics
As an integral component of the mammalian cell cytoskeleton, actin is involved in multiple physiological functions, including cell growth, motility, trafficking, and division. These basic functions require remodeling of the actin cytoskeleton, as well as extension and withdrawal of actin-based cellular neurites (e.g., lamellipodia, filopodia), all of which rely upon rapid dynamic cycling between filamentous actin (F-actin) and monomer actin (G-actin)1. Correspondingly, dysfunctional actin cytoskeletal dynamics are a pathophysiological feature of many human diseases, with perhaps cancer being the prototypical example. For these reasons, actin is a theoretically attractive anti-cancer therapeutic target. However, in practice, actin has proven to be a poor target because of toxic side effects due in large part to the inability of therapeutics to distinguish between actin isoforms2,3. Thus, the actin in cancerous and healthy cells are affected by cancer therapies acting directly upon actin, leading to toxic side effects in different organ systems (e.g., heart and diaphragm)2,3. Recently, drug discovery has shifted from actin to actin-associated structural proteins such as the Arp2/3 complex and tropomyosins (Tpms) as these proteins offer multiple isoforms for selective targeting and the opportunity to avoid toxic side effects2-4 (Fig. 1).
Developing drugs that regulate actin dynamics by targeting structural proteins is not without challenges. Necessary structural knowledge of target proteins is obtained through functional domain mapping and identification of possible drug binding pockets prior to the start of rational drug design3. In the case of actin-associated regulatory proteins such as Arp2/3 and Tpms, these data are lacking (Fig. 1). In the absence of complete structural data, researchers screen existing compound libraries against target proteins, searching for drugs that inhibit function3. An attribute of a putative therapeutic target is the existence of multiple isoforms which offer the possibility of selective targeting, especially if only certain isoforms are associated with disease. Moreover, structural isoforms offer more druggable binding pockets which means that different drugs can regulate the same protein, potentially with different mechanisms of action. To take advantage of existing small molecule compound libraries and deal with the lack of structural knowledge of candidate actin-binding proteins, one technique that is employed is phenotypic profiling3. This screening protocol utilizes cellular imaging to measure how compounds affect multiple, complex cellular architectures, including cell size, morphology, and in the case of actin, specific details of the filaments’ attributes (number, length, shape, subcellular distribution, signal intensity) by computational and automated image analyses to identify compounds producing relevant biological effects3,5-7. Furthermore, when a relevant compound’s mechanism of action is known, this information is used to both further evaluate this compound and predict the mechanisms of action of other compounds that produce similar phenotypes3,7.
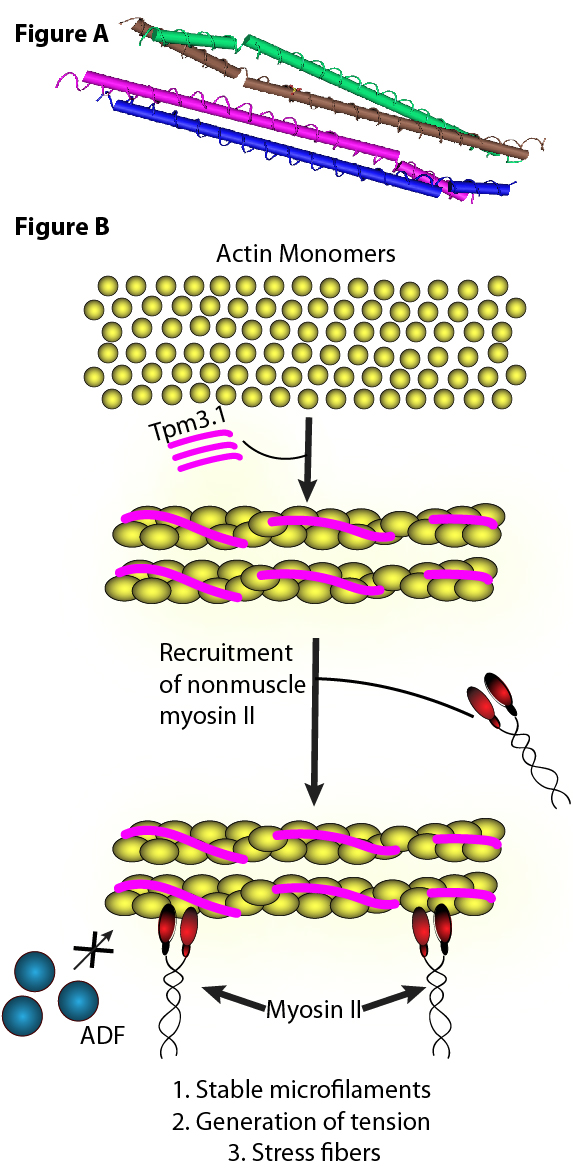
Figure A. Tropomyosin isoform 3.1 (Tpm3.1) is a cancer-associatedactin-binding protein.(Top) Crystal structure of an N-terminal fragment of Tpm3.1. PDB ID: 6OTN.
Figure B. (Bottom) Tpm3.1 regulates how F-actin interacts with non-muscle myosin II and ADF/cofilin.Adapted from reference (12).
The Arp2/3 complex and Tpms are promising therapeutic targets in cancer drug discovery programs3,8-12. Arp2/3 activity can be inhibited by different drugs through different mechanisms of actions11. Tpm isoforms differentially regulate how susceptible F-actin is to actin-modulating drugs3,12. Tpms consist of over 40 isoforms and display individual spatial and temporal localization patterns during cell/tissue development12. The actin cytoskeleton consists of both co-polymers of actin and Tpms and polymers of Tpm-free actin13; however, a recent study reports that Tpms coat the majority of F-actin in human cells14. Tpms determine the functional capability of actin filaments in an isoform-dependent manner15-21 (Fig. 1). One important characteristic of a protein that qualifies it for further study as a therapeutic target is its specific modulation in activity/expression in cancer cell lines. In the case of Tpms, isoform 3.1 is significantly and selectively up-regulated in multiple cancer cell lines; thus, it is a disease-associated isoform that can be specifically targeted2,14,22,23 (Fig. 1). Importantly, targeting Tpm3.1 also avoids toxic side effects as muscle Tpms do not express this isoform22,24,25. Currently under investigation are three structurally unique, small molecule compounds that selectively target Tpm3.1 via distinct molecular interactions within Tpm3.1’s binding pocket3,22,24,25.
Summary
Investigating actin-associated structural proteins as novel anti-cancer therapeutic targets offers great potential for the development of novel and specific cancer therapies, but there are several caveats and methodological considerations when using phenotypic screening. For instance, 1. candidate cell lines must possess a highly conserved actin cytoskeleton; 2. any actin cytoskeleton stains must be compatible with robotic manipulation; 3. different actin phenotypes must be detectable with the specific screening set-up used and multiple parameters must be measurable (e.g., filament number and length, cell morphology and size, subcellular distribution of F-actin and its distinct structural forms); and 4. actin dynamics within cardiac and skeletal muscle cells must not be affected3. To assist researchers in screening compounds for their interactions with actin and associated proteins, Cytoskeleton offers a variety of purified actin proteins and actin-associated proteins, actin activity and binding assay kits, and live cell imaging probes for F-actin.
References
- Pollard T.D. 2016. What we know and do not know about actin. In: Jockusch B. (eds.) The Actin Cytoskeleton. Handbook of Experimental Pharmacology. vol 235. Springer, Cham.
- Bonello T.T. et al. 2009. New approaches to targeting the actin cytoskeleton for chemotherapy.Future Med. Chem. 1, 1311-1331.
- Bryce N.S. et al. 2019. Chemical biology approaches targeting the actin cytoskeleton through phenotypic screening. Curr. Opin. Chem. Biol. 51, 40-47.
- Creed S.J. et al. 2008. Tropomyosin isoforms define distinct microfilament populations with different drug susceptibility. Eur. J. Cell Biol. 87, 709-720.
- Feng Y. et al. 2009. Multi-parameter phenotypic profiling: using cellular effects to characterize small-molecule compounds. Nat. Rev. Drug Discov. 8, 567-578.
- Eliceiri K.W. et al. 2012. Biological imaging software tools. Nat. Methods. 9, 697-710.
- Caicedo J.C. et al. 2017. Data-analysis strategies for image-based cell profiling. Nat. Methods. 14, 849-863.
- Jay P. et al. 2000. ARP3beta, the gene encoding a new human actin-related protein, is alternatively spliced and predominantly expressed in brain neuronal cells.Eur. J. Biochem. 267, 2921-2928.
- Millard T.H. et al. 2003. Identification and characterisation of a novel human isoform of Arp2/3 complex subunit p16-ARC/ARPC5.Cell Motil. Cytoskeleton. 54, 81-90.
- Abella J.V. et al. 2016. Isoform diversity in the Arp2/3 complex determines actin filament dynamics. Nat. Cell Biol. 18, 76-86.
- Nolen B.J. et al. 2009. Characterization of two classes of small molecule inhibitors of Arp2/3 complex. Nature. 460, 1031–1034.
- Gunning P.W. et al. 2015. Tropomyosin - master regulator of actin filament function in the cytoskeleton. J. Cell Sci. 128, 2965-2974.
- Lazarides E. 1976. Two general classes of cytoplasmic actin filaments in tissue culture cells: the role of tropomyosin. J. Supramol. Struct. 5, 531–563.
- Meiring J.C.M. et al 2018. Co-polymers of actin and tropomyosin account for a major fraction of the human actin cytoskeleton. Curr. Biol. 28, 2331-2337.
- Bryce N.S. et al. 2003. Specification of actin filament function and molecular composition by tropomyosin isoforms. Mol. Biol. Cell14, 1002–1016.
- Tojkander S. et al. 2011. A molecular pathway for myosin II recruitment to stress fibers. Curr. Biol. 21, 539–550.
- Creed S.J. et al. 2011. Tropomyosin isoform 3 promotes the formation of filopodia by regulating the recruitment of actin-binding proteins to actin filaments. Exp. Cell Res. 317, 249–261.
- Janco M. et al. 2016. The impact of tropomyosins on actin filament assembly is isoform specific. Bioarchitecture6, 61–75.
- Brayford S. et al. 2016. Tropomyosin promotes lamellipodial persistence by collaborating with Arp2/3 at the leading edge. Curr. Biol.26, 1312–1318.
- Gateva G. et al. 2017. Tropomyosin isoforms specify functionally distinct actin filament populations in vitro. Curr. Biol. 27, 705–713.
- Pathan-Chhatbar S. et al. 2018. Three mammalian tropomyosin isoforms have different regulatory effects on nonmuscle myosin-2B and filamentous b-actin in vitro. J. Biol. Chem. 293, 863–875.
- Stehn J.R. et al. 2013. A novel class of anticancer compounds targets the actin cytoskeleton in tumor cells. Cancer Res.73, 5169-5182.
- Coombes J.D. et al 2015. Ras transformation overrides a proliferation defect induced by Tpm3.1 knockout. Cell. Mol. Biol. Lett. 20, 626-646.
- Currier M.A. et al. 2017. Identification of cancer-targeted tropomyosin inhibitors and their synergy with microtubule drugs. Mol. Cancer Ther. 16, 1555-1565.
- Kee A.J. et al. 2018. On-target action of anti-tropomyosin drugs regulates glucose metabolism. Sci. Rep. 8, 4604.
Related Products
Fluorescent Tubulins
Tubulin protein (fluorescent AMCA): porcine brain (Cat. TL440M)
Tubulin protein (fluorescent HiLyte 488): porcine brain (Cat. # TL488M)
Tubulin protein (rhodamine): porcine brain (Cat. # TL590M)
Tubulin protein (X-rhodamine): bovine brain (Cat. # TL620M)
Tubulin protein (fluorescent HiLyte 647): porcine brain (Cat. # TL670M)
Signal Seeker™ Kits
Signal-Seeker™ Phosphotyrosine Detection Kit (30 assay) (Cat. # BK160)
Signal-Seeker™ Phosphotyrosine Detection Kit (10 assay) (Cat. # BK160-S)
Signal-Seeker™ Ubiquitination Detection Kit (30 assay) (Cat. # BK161)
Signal-Seeker™ Ubiquitination Detection Kit (10 assay) (Cat. # BK161-S)
Signal-Seeker™ SUMOylation 2/3 Detection Kit (30 assay) (Cat. # BK162)
Signal-Seeker™ SUMOylation 2/3 Detection Kit (10 assay) (Cat. # BK162-S)
Signal-Seeker™ Acetyl-Lysine Detection Kit (30 assay) (Cat. # BK163)
Signal-Seeker™ Acetyl-Lysine Detection Kit (10 assay) (Cat. # BK163-S)
NEW Signal-Seeker™ SUMOylation 1 Detection Kit (30 assay) (Cat. # BK165)
NEW Signal-Seeker™ SUMOylation 1 Detection Kit (10 assay) (Cat. # BK165-S)
Tubulin Kits
Tubulin polymerization HTS assay using >97% pure tubulin, OD based - Porcine (Cat. # BK004P)
Tubulin polymerization assay using >99% pure tubulin, OD based - Porcine (Cat. # BK006P)
Tubulin polymerization assay using >99% pure tubulin, fluorescence based (Cat. # BK011P)
Microtubule Binding Protein Spin-Down Assay Biochem Kit (Cat. # BK029)
Microtubule/Tubulin In Vivo Assay Biochem Kit (Cat. # BK038)
Live Cell Imaging Products

