Profilin: Multi-functional Roles of an Actin Binding Protein
The profilin (Pfn) family of proteins were originally characterized and studied as regulators of actin polymerization. First identified in 19761,2, these small (14-17 kDa) proteins exist as four isoforms in humans (Pfn1-4)3. Pfn1 is expressed in most cell types, Pfn2 is primarily localized to the brain, and Pfn3 and Pfn4 are localized to the testes4. Pfn expression is essential for the embryonic development of mice5,6. However, forty years later, the role of Pfn proteins in regulating actin polymerization, activating intracellular signal transduction pathways via binding to polyphosphoinositides3,6, regulating microtubule end turnover7, binding ligands via poly-L-proline domains3,6, and potentially suppressing tumorigenicity is still being investigated8-10. This newsletter describes the different biological interactions that Pfns are involved in and how these interactions affect actin polymerization.
Profilins and Actin Polymerization
Cellular processes such as trafficking, motility, division, and growth require remodeling of the actin cytoskeleton. Pfns regulate actin polymerization and can both inhibit and promote actin polymerization11. Pfns bind to monomeric G-actin in a 1:1 ratio with a binding affinity of 0.1 µM, effectively sequestering the G-actin from incorporation into growing filaments (Fig. 1). Notably, the intracellular concentration of Pfns has been estimated to be 10-80 µM, which is not sufficient to maintain the high concentrations of G-actin found in the cell12-14. Pfns can also bind the barbed end of F-actin, albeit with a reduced affinity (25 µM)15-17. In addition, Pfn1 catalyzes the nucleotide exchange of ADP for ATP on G-actin by 1000-fold, which resupplies the pool of ATP-G-actin for barbed end growth6,18,19. Indeed, when the barbed ends of an actin filament are exposed, the ATP-actin:Pfn complex promotes filament elongation by loading ATP-actin onto the barbed end of the actin filament, followed by disassociation of the bound Pfn protein6,17. However, in the presence of end-capping proteins bound to the barbed ends of actin polymers, it has been observed in vitro that Pfn acts as an actin sequestering protein that prevents the formation of actin filaments and promotes F-actin depolymerization11,20. Depolymerization of F-actin was also observed in vivo upon microinjection of Pfn into either Swiss 3T3 fibroblasts or rat kidney cells21.
While the exact molecular mechanisms by which Pfns promote and inhibit actin polymerization are complex, Pfns appear to act as a gatekeeper in regulating the activities of the actin nucleation-promoting factors, formins and Arp2/3 complex22. Formins direct the formation of linear, unbranched filaments, while the Arp2/3 complex and the VCA domain of WASP stimulate the nucleation of branched actin filaments. Recent studies show that the Pfns favor formin-mediated actin polymerization and inhibit Arp2/3-mediated actin polymerization22,23. While the exact molecular mechanisms underlying Pfn-mediated inhibition of Arp2/3 activities are unknown, it is speculated that Pfns provide actin monomers for formin-mediated activity, while directly or indirectly competing with the VCA domain of WASP for actin monomer binding23.
Profilins and Phosphoinositide Binding
Polyphosphoinositides modulate actin polymerization, at least partially through an interaction with Pfns. Pfns interact with both phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2) and phosphatidylinositol (3,4,5)-trisphosphate (PI(3,4,5)P3)24-26. Binding of PI(4,5)P2 to Pfn is believed to prevent Pfn from binding G-actin, thereby increasing the pool of G-actin available for Arp2/3-mediated branched filament formation. Alternatively, an increase in Pfn following PI(4,5)P2 hydrolysis by phosphorylated phospholipase C could decrease the G-actin pool and promote formin-mediated actin polymerization27. To date, phosphoinositide-mediated control of Pfn-regulated actin assembly has only been observed in vitro.
Profilins and Poly-L-Proline
Pfns also recognize poly-L-proline stretches via their N- and C-terminal helices, domains distinct from the actin binding domain (Fig. 1). The capacity of Pfns to bind poly-L-proline enables binding to over 50 different ligands in multiple organisms6. While several Pfn binding partners are involved in actin regulation, other newly discovered partners are involved in endocytosis, nuclear export, and Rac/Rho effector protein signaling6. These vastly different binding partners suggest that Pfns may be regulated or involved in other non-actin-related pathways. The physiological relevance of these interactions is not well understood and awaits elucidation.
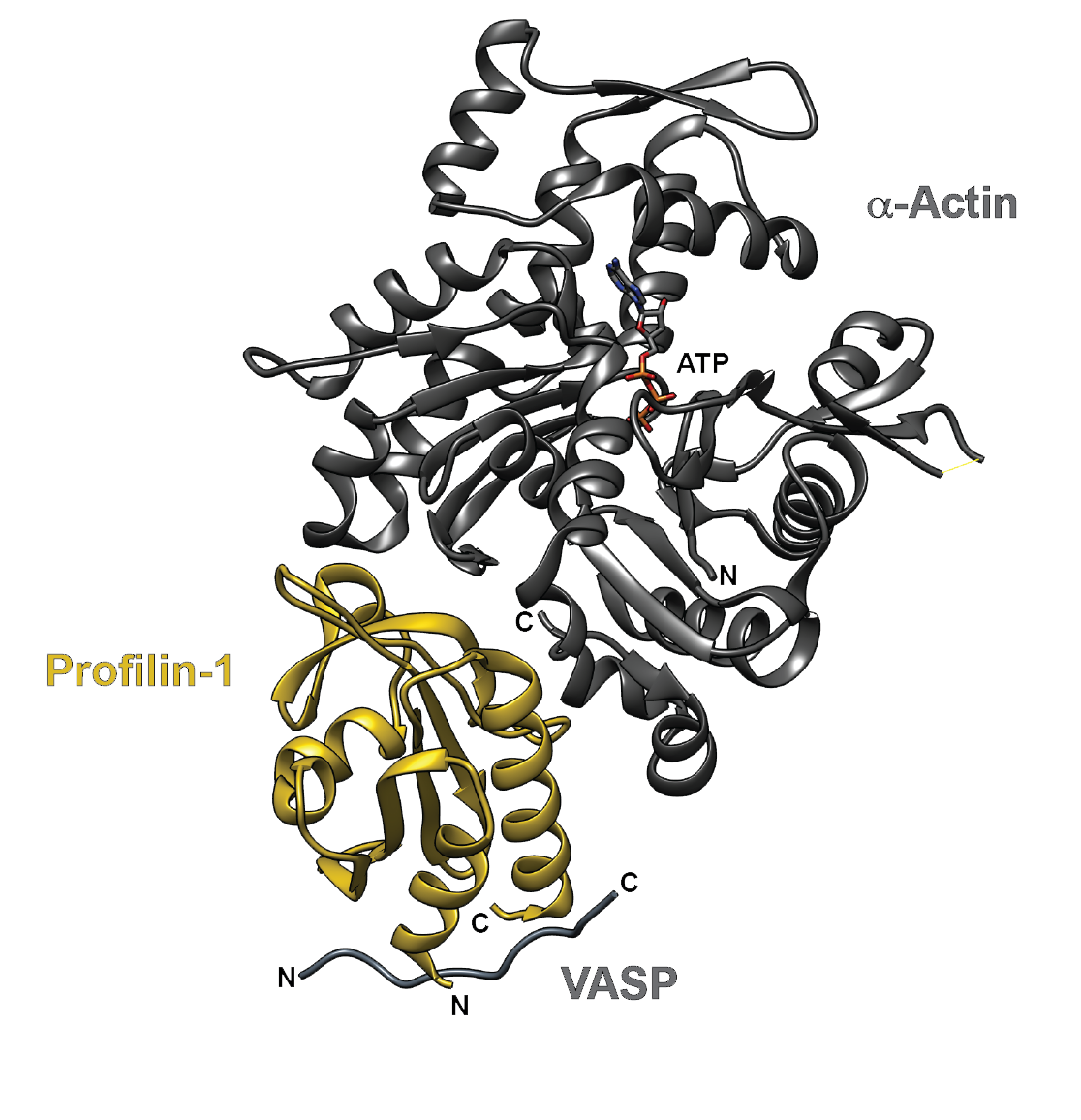
Figure 1. The crystal structure of human profilin-1 (yellow) bound on opposite sides to α-actin (grey) and the peptide from the poly-proline site of human VASP. Image was created with UCSF Chimera30 (PDB 2PAV).
Profilins and Cancer
Dynamic changes in the actin cytoskeleton occur during metastasis and cancer cell invasion. Several breast cancer cell lines with higher levels of expression for Pfn1 have reduced tumorigenicity, motility, and invasiveness8,28. Reduced binding of Pfn to actin in one breast cancer cell line exhibited reduced suppression of tumorgencity29. Pfn overexpression reduces the invasiveness of some breast cancer cells lines and orthotopic cancer systems8,28. These results suggest that the control of cytoskeleton dynamics by Pfns can play an important role in inhibiting cancer cell proliferation and migration and warrant further investigation.
Summary
Despite decades of research, the functionality and binding partners of the actin binding protein Pfn are not completely understood. Indeed, Pfns exert a complex regulation on actin, both able to inhibit and promote actin filament formation. Perhaps even more intriguing are recent reports that Pfn’s functions go far beyond regulating actin cytoskeletal dynamics to include roles in controlling microtubule dynamics, binding proline domain-containing ligands, and activating a multitude of intracellular signaling pathways via binding to polyphosphoinositides. Pfns also offer potential as a drug target in anti-cancer therapies. To help researchers unravel Pfn’s various functions and binding partners, Cytoskeleton, Inc. offers purified actin and actin binding proteins, including profilin, Acti-stain phalloidins, functional assay kits, and F-actin live cell imaging probes.
Related Products & Resources
Actin Products
Profilin 1 (recombinant human no tag) (Cat. # PR02)
Actin Protein (>95% pure): rabbit skeletal muscle (Cat. # AKL95)
Actin Protein (>99% pure): rabbit skeletal muscle (Cat. # AKL99)
Spirochrome SiR-Actin Kit (Cat. # CY-SC001)
Spirochrome SiR700-Actin Kit (Cat. # CY-SC013)
Acti-stain™ 488 Phalloidin (Cat. # PHDG1)
Acti-stain™ 555 Phalloidin (Cat. # PHDH1)
Phalloidin (rhodamine) (Cat. # PHDR1)
Actin Biochem Kits
Actin Binding Protein Spin-Down Assay Biochem Kit: rabbit skeletal muscle actin (Cat. # BK001)
Actin Binding Protein Spin-Down Assay Biochem Kit: human platelet actin (Cat. # BK013)
Actin Polymerization Biochem Kit (fluorescence format): rabbit skeletal muscle actin (Cat. # BK003)
G-Actin/F-actin In Vivo Assay Biochem Kit (Cat. # BK037)
References
- Carlsson L. et al. 1976. Crystallization of a non-muscle actin. J. Mol. Biol. 105, 353–366.
- Carlsson L. et al. 1977. Actin polymerizability is influenced by profilin, a low molecular weight protein in non-muscle cells. J. Mol. Biol. 115, 465–483.
- Krishnan K. and Moens P.D.J. 2009. Structure and functions of profilins. Biophys. Rev. 1, 71–81.
- Aspenström P. 2010. Formin-binding proteins: modulators of formin-dependent actin polymerization. Biochim. Biophys. Acta. 1803, 174–182.
- Witke W. et al. 2001. Profilin I is essential for cell survival and cell division in early mouse development. Proc. Natl. Acad. Sci. U.S.A. 98, 3832–3836.
- Witke W. 2004. The role of profilin complexes in cell motility and other cellular processes. Trends Cell Biol. 14, 461–469.
- Nejedla M. et al. 2016. Profilin connects actin assembly with microtubule dynamics. Mol. Biol. Cell. 27, 2381–2393.
- Janke J. et al. 2000. Suppression of tumorigenicity in breast cancer cells by the microfilament protein profilin 1. J. Exp. Med. 191, 1675–1686.
- Adami G.R. et al. 2017. A loss of profilin-1 in late-stage oral squamous cell carcinoma. J. Oral Pathol. Med. 46, 489–495.
- Schoppmeyer R. et al. 2017. Human profilin 1 is a negative regulator of CTL mediated cell-killing and migration. Eur. J. Immunol. 47, 1562–1572.
- Yarmola E.G. and Bubb M.R. 2006. Profilin: emerging concepts and lingering misconceptions. Trends Biochem. Sci. 31, 197–205.
- Southwick F.S. and Young C.L. 1990. The actin released from profilin--actin complexes is insufficient to account for the increase in F-actin in chemoattractant-stimulated polymorphonuclear leukocytes. J. Cell Biol. 110, 1965–1973.
- Moldovan N.I. et al. 1997. Regulation of endothelial cell adhesion by profilin. Curr. Biol. 7, 24–30.
- Pernier J. et al. 2016. Profilin interaction with actin filament barbed end controls dynamic instability, capping, branching, and motility. Dev. Cell. 36, 201–214.
- Perelroizen I. et al. 1994. Interaction of profilin with G-actin and poly(L-proline). Biochemistry. 33, 8472-8478.
- Vinson V.K. et al. 1998. Interactions of Acanthamoeba profilin with actin and nucleotides bound to actin. Biochemistry. 37, 10871–10880.
- Shekhar S. et al. 2016. Regulators of actin filament barbed ends at a glance. J. Cell Sci. 129, 1085–1091.
- Tilney L.G. et al. 1983. Actin from Thyone sperm assembles on only one end of an actin filament: a behavior regulated by profilin. J. Cell Biol. 97, 112–124.
- Pring M. et al. 1992. Profilin-actin complexes directly elongate actin filaments at the barbed end. Biochemistry. 31, 1827–1836.
- Bubb M.R. et al. 2003. Depolymerization of actin filaments by profilin. Effects of profilin on capping protein function. J. Biol. Chem. 278, 24629–24635.
- Hájková L. et al. 1997. Characterization of a mutant profilin with reduced actin-binding capacity: effects in vitro and in vivo. Exp. Cell Res. 234, 66–77.
- Rotty J.D. et al. 2015. Profilin-1 serves as a gatekeeper for actin assembly by Arp2/3-dependent and -independent pathways. Dev. Cell. 32, 54–67.
- Suarez C. et al. 2015. Profilin regulates F-actin network homeostasis by favoring formin over Arp2/3 complex. Dev. Cell. 32, 43–53.
- Lassing I. and Lindberg U. 1985. Specific interaction between phosphatidylinositol 4,5-bisphosphate and profilactin. Nature. 314, 472–474.
- Fedorov A.A. et al. 1994. X-ray structures of isoforms of the actin-binding protein profilin that differ in their affinity for phosphatidylinositol phosphates. Proc. Natl. Acad. Sci. U.S.A. 91, 8636–8640.
- Chaudhary A. et al. 1998. Probing the phosphoinositide 4,5-bisphosphate binding site of human profilin I. Chem. Biol. 5, 273–281.
- Bezanilla M. et al. 2015. Cytoskeletal dynamics: a view from the membrane. J. Cell Biol. 209, 329–337.
- Zou L. et al. 2007. Profilin-1 is a negative regulator of mammary carcinoma aggressiveness. Br. J. Cancer. 97, 1361–1371.
- Wittenmayer N. et al. 2004. Tumor suppressor activity of profilin requires a functional actin binding site. Mol. Biol. Cell. 15, 1600–1608.
- Pettersen E.F. et al. 2004. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605-1612.

