RhoA / Rac1 / Cdc42 Activation Assay Combo Biochem Kit (bead pull-down format) - 3 x 10 assays
Product Uses Include
- The combo kit is an ideal choice when your experimental observations suggest that one or more Rho family proteins are activated.
- Provides 10 assays each for detection of activated RhoA, Rac1, and Cdc42.
- Includes an extensive array of reagents (see kit contents below).
- Most cost effective way to obtain a Rho family activation profile.
Assay Principle
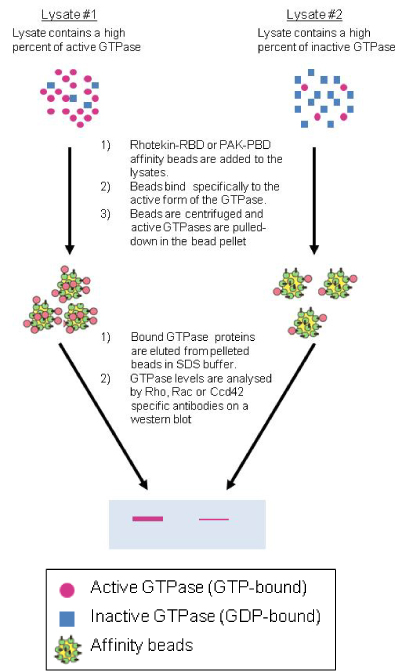
The principle of the assay is shown schematically below (Figure 1). The assay is based upon the fact that a Rho family effector protein is known to bind preferentially to the active (GTP-bound) form of its target GTPase (1). In the case of RhoA activation, the rhotekin-RBD effector domain is used to make the affinity beads (2). In the case of Cdc42 and Rac1 activation, the PAK-PBD effector domain is used for affinity beads (3). Example Western blot results and detailed protocols can be viewed by clicking the documents tab above and downloading the pdf datasheet.
Kit contents
The kit contains sufficient materials for 10 assays each for RhoA, Rac1, and Cdc42 (depending on activation levels in cells), including reagents for positive and negative controls. The following components are included:
- GST-tagged Rhotekin-RBD protein on colored agarose beads (Part # RT02-S)
- GST-tagged PAK-PBD protein on colored agarose beads (Part # PAK02-S)
- RhoA-specific monoclonal antibody, 50 ug (Cat. # ARH03)
- Rac1-specific monoclonal antibody, 50 ug (Cat. # ARC03)
- Cdc42 monoclonal antibody, 50 ug (Cat. # ACD03)
- His-tagged RhoA protein (Part # RHWT)
- His-tagged Rac1 protein (Part # RCWT)
- His-tagged Cdc42 protein (Part # CDWT)
- GTPγS: (non-hydrolyzable GTP analog) (Cat. # BS01)
- GDP (Part # GDP01)
- Cell lysis Buffer (Part # CLB01)
- Wash Buffer (Part # WB01-S)
- Loading Buffer (Part # LB01)
- STOP Buffer (Part # STP01)
- DMSO (Part # DMSO)
- Protease inhibitor cocktail (Cat. # PIC02)
- Manual with detailed protocols and extensive troubleshooting guide
Equipment needed
- SDS-PAGE minigel system and Western blotting transfer apparatus
References
- Aspenstrom, P. 1999. Effectors for the Rho GTPases. Curr. Opin. In Cell Biol. 11, 95-102.
- Ren, X.D. et al. 1999. Regulation of the small GTP-binding protein Rho by cell adhesion and the cytoskeleton. EMBO J. 18, 578-585.
- Benard, V. et al. 1999. Characterization of Rac and Cdc42 activation in chemoattractant-stimulated human neutrophils using a novel assay for active GTPases. J. Biol. Chem. 274: 13198-13204 .
Please check out the new versions of Activation Assays and associated products:
G-LISA® Activation Assays:
Cdc42 G-LISA® Activation Assay, colorimetric format (Cat.# BK127)
Rac1 G-LISA® Activation Assay, luminescence format (Cat.# BK126)
Rac1,2,3 G-LISA® Activation Assay, colorimetric format (Cat.# BK125)
RhoA G-LISA® Activation Assay, colorimetric format (Cat.# BK124)
RhoA G-LISA® Activation Assay, luminescence format (Cat.# BK121)
Large Pull-down Activation Assays:
Cdc42 Activation Assay Biochem Kit, bead pull-down format (Cat.# BK034)
Rac1 Activation Assay Biochem Kit, bead pull-down format (Cat.# BK035)
RhoA Activation Assay Biochem Kit, bead pull-down format (Cat.# BK036)
Associated Products:
Anti-Cdc42 monoclonal antibody (Cat.# ACD03)
Anti-Rac1 monoclonal antibody (Cat.# ARC03)
Anti-RhoA monoclonal antibody (Cat.# ARH03)
GST-tagged Rhotekin-RBD protein on colored agarose beads (Cat. # RT02)
GST-tagged PAK-PBD protein on colored agarose beads (Cat. # PAK02)
His-tagged RhoA protein (Cat. # RH01)
His-tagged Rac1 protein (Cat. # RC01)
His-tagged Cdc42 protein (Cat. # CD01)
For product Datasheets and MSDSs please click on the PDF links below. For additional information, click on the FAQs tab above or contact our Technical Support department at tservice@cytoskeleton.com
Question 1: I have high background and/or multiple bands on my western blot. How can I fix this?
Answer 1: There are multiple causes of high background and/or multiple bands. Some suggestions to improve background signal include:
- When blotting use 70v for 45min only as the small G-proteins are very mobile.
- Fully remove SDS from the gel by using a non-SDS containing buffer for transfer and performing a full 15 min gel wash step in the transfer buffer before blotting.
- Dry the PVDF membrane for 30 min after transfer and before blocking (not necessary for nitrocellulose)
- Making sure that the TBST contains 10 mM Tris, 0.05% Tween 20 and 150 mM NaCl.
- Incubating with the primary antibody overnight at 4°C and using the appropriate ECL detection system.
Question 2: How much of the beads should I use for my pull-down experiments?
Answer 2: The beads conjugated to the respective effector protein that recognizes the active form of each GTPase will bind to the GDP-bound GTPase with a much lower affinity than the GTP-bound GTPase. If too many beads are added to the pull-down assay there will be significant binding to inactive (GDP-bound) GTPases. The result of this will be an underestimation of GTPase activation. For this reason, we highly recommend performing a bead titration to determine optimal conditions for any given GTPase activation or inactivation assay. Once optimal conditions have been established, bead titrations should no longer be necessary.
Question 3: How can I test whether the beads are working properly?
Answer 3: A standard biological assay for the beads consists of a GTPase protein pull-down from cells loaded with either GTPγS (Cat. # BS01) or GDP. Here are guidelines to follow (see Cat. # BK030 manual for more details):
Positive Cellular Protein Control:
Total cell lysate (300 – 800 μg) should be loaded with GTPγS as a positive control for the pull-down assay. The following reaction details how to load endogenous GTPase with the nonhydrolysable GTP analog (GTPγS). This is an excellent substrate for the beads and should result in a strong positive signal in a pull-down assay.
a) Perform GTP loading on 300 – 800 μg of cell lysate (0.5 mg/ml protein concentration) by adding 1/10th volume of Loading Buffer.
b) Immediately add 1/100th volume of GTPγS (200 μM final concentration). Under these conditions, 5 - 10% of the GTPase protein will load with non-hydrolysable GTPγS and will be “pulled-down” with the beads in the assay.
c) Incubate the control sample at 30°C for 15 min with gentle rotation.
d) Stop the reaction by transferring the tube to 4°C and adding 1/10th volume of STOP Buffer.
e) Use this sample immediately in a pull-down assay.
Negative Cellular Protein Control:
This reaction should be performed in an identical manner to the Positive Control reaction except that 1/100th volume of GDP (1 mM final concentration) should be added to the reaction in place of the GTPγS. Loading endogenous GTPase with GDP will inactivate the GTPase and this complex will bind very poorly to the beads.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com