Rac1,2,3 G-LISA Activation Assay (Colorimetric format) 96 assays
Product Uses Include
- Rac signaling pathway studies
- Rac activation assays with primary cells
- Studies of Rac activators and inactivators
- Rac activation assays with limited material
- High throughput screens for Rac activation
Introduction
The G-LISA series of Small G-Protein Activation Assays are ELISA based assays with which you can measure the GTP form of small G-proteins from lysates of cells or tissues and all in less than 3 h. For a more detailed introduction on G-LISA™ assays and a listing of other available G-LISA™ kits, see our main G-LISA™ page. The Rac1,2,3 G-LISA™ Activation Assay measures the entire level of GTP-loaded Rac1,2 and 3 protein in cell lysates, this is in contrast to Cat.# BK126 which measures only Rac1 activation levels. The level of activation is measured with absorbance at 490nm. For a kit to measure RhoA activation please check webpage BK124.
The Rac G-LISA Activation Assay is very sensitive and has excellent accuracy for duplicate samples. See G-LISA™ FAQs tab on our G-LISA™ page for more details.
Kit contents
The kit contains sufficient reagents to perform 96 Rac activation assays. Since the Rac-GTP affinity wells are supplied as strips and the strips can be broken into smaller pieces, each kit can be used for anywhere from one to multiple assays. The following components are included in the kit:
- 96 Rac-GTP affinity wells (divisible into 12 strips of 8 wells each)
- Lysis buffer
- Binding buffer
- Antigen presenting buffer
- Wash buffer
- Antibody dilution buffer
- Anti-Rac antibody
- HRP-labeled secondary antibody
- Positive control Rac protein
- Protease inhibitor cocktail (Cat. # PIC02)
- Absorbance detection reagents
- Precision Red™ Advanced protein assay reagent (Cat. # ADV02)
- Manual with detailed protocols and extensive troubleshooting guide
Equipment needed
- 96-well plate spectrophotometer capable of reading 490 nm wavelength
- Multichannel or multidispensing pipettor
- Orbital microplate shaker capable of at least 200 rpm shaking (400 rpm is optimal)
Example results
Serum starved Swiss 3T3 cells were stimulated with the Rac activating compound EGF and Rac activation was measured with the G-LISA method (Fig 1 and 2).
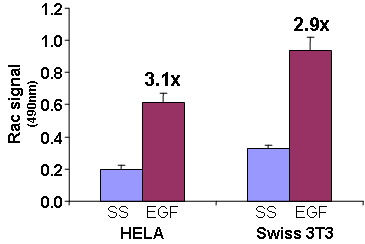
Figure 1. Rac activation by EGF measured by G-LISA™ kit BK125. Swiss 3T3 (mouse) cells were serum starved for 24 h and treated with EGF (Cal; 10ng/ml for 3 min) or buffer only (SS). 10 µg of cell lysates were subjected to the G-LISA™ assay. Absorbance was read at 490 nm.
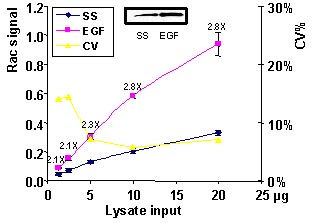
Figure 2. Rac activation by EGF measured by G-LISA™. Swiss 3T3 cells were serum starved (SS) for 24 h and treated with EGF (10 ng/ml for 2 min). 20, 10, 5, 2.5, 1.25 µg of cell lysates were subjected to the G-LISA™ assay. Absorbance was read at 490 nm. 500 µg of the same lysates were subjected to the traditional PAK pull-down assay (shown in inset, Cat.# BK035).
G-LISA™ Products:
Cdc42 G-LISA™ Activation Assay, colorimetric format (Cat.# BK127)
Rac1 G-LISA™ Activation Assay, luminescence format (Cat.# BK126)
RhoA G-LISA™ Activation Assay, colorimetric format (Cat.# BK124)
RhoA G-LISA™ Activation Assay, luminescence format (Cat.# BK121)
Associated Products:
Anti-Cdc42 monoclonal antibody (Cat.# ACD03)
Anti-Rac1 monoclonal antibody (Cat.# ARC03)
Anti-RhoA monoclonal antibody (Cat.# ARH03)
For product Datasheets and MSDSs please click on the PDF links below.
 G-LISA Activation Assay Technical Guide download here
G-LISA Activation Assay Technical Guide download here G-LISA Data Analysis (Absorbance) Excel Template download here.
G-LISA Data Analysis (Absorbance) Excel Template download here.
Question 1: Can I detect isoforms other than RhoA, Rac1,2,3 or RalA with these G-LISA activation assays?
Answer 1: Yes, the RhoA G-LISA (Cat. # BK124), Rac1,2,3 G-LISA (Cat. # BK125) and RalA G-LISA (Cat. # BK129) can be used to detect RhoB or RhoC, Rac 2 or Rac3 or RalB, respectively. To specifically detect Rac1, please see our Rac1 G-LISA activation assay (Cat. # BK128). The capture proteins that the wells have been coated with bind all of the isoforms of the respective GTPase. The specificity of signal is conferred by the specificity of the monoclonal primary antibody utilized. Use of an isoform-specific monoclonal antibody allows detection of other Rho family isoforms. Please see this citation for an example of this modified procedure (Hall et al., 2008. Type I Collagen Receptor (α2β1) Signaling Promotes Prostate Cancer Invasion through RhoC GTPase. Neoplasia. 10, 797–803).
Basically the researcher would test their specific monoclonal antibody in a western blot first to prove specificity to the alternative isoform of interest. For example, load RhoA and C for negative controls when testing a RhoB monoclonal antibody. Then the researcher would use 1:50, 1:200 and 1:500 dilutions of their monoclonal antibody on duplicate cell extracts of activated and control state samples. The researcher would then choose the dilution of monoclonal antibody which gave them the highest ratio of activated:control state.
A simple activated/control state pair of extracts can be made by growing cells to 50% confluence in serum containing media, washing twice with PBS, preparing lysate and aliquoting and freezing samples in liquid nitrogen. With one aliquot, defrost and let stand at room temperature for 60 min to degrade the activated signal to a low basal signal, which will be the control state. The untreated sample (2nd aliquot) will be considered “activated” which most serum grown cells are.
Question 2: How many cell culture plates can I process at one time during the lysis step?
Answer 2: We recommend that from the point at you add lysis buffer to the plate on ice to aliquoting and snap-freezing the lysate samples in liquid nitrogen, no more than 10 min are allowed to elapse. After 10 min on ice, we find that GTP bound to GTPases (activated GTPases) undergoes rapid hydrolysis. Rapid processing at 4°C is essential for accurate and reproducible results. The following guidelines are useful for rapid lysis of cells.
Washing
a. Retrieve culture dish from incubator, immediately aspirate out all of the media and place firmly on ice.
b. Immediately rinse cells with an appropriate volume of ice cold PBS (for Cdc42 activation, skip this step and simply aspirate the media) to remove serum proteins.
c. Aspirate off all residual PBS buffer. This is essential so that the Lysis Buffer is not diluted. Correct aspiration requires that the culture dish is placed at a steep angle on ice for 1 min to allow excess PBS to collect in the vessel for complete removal. As noted, the time period between cell lysis and addition of lysates to the wells is critically important. Take the following precautions:
1. Work quickly.
2. Keeping solutions and lysates embedded in ice so that the temperature is below 4°C. This helps to minimize changes in signal over time.
3. We strongly recommend that cell lysates be immediately frozen after harvest and clarification. A sample of at least 20 μl should be kept on ice for protein concentration measurement. The lysates must be snap frozen in liquid nitrogen and stored at -70°C. Lysates should be stored at -70°C for no longer than 30 days.
4. Thawing of cell lysates prior to use in the G-LISA assay should be in a room temperature water bath, followed by rapid transfer to ice and immediate use in the assay.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com.








