Rac1 G-LISA Activation Assay (Luminescence format) - 96 assays
Product Uses Include
- Rac1 signaling pathway studies
- Rac1 activation assays with primary cells
- Studies of Rac1 activators and inactivators
- Rac1 activation assays with limited material
- High throughput screens for Rac1 activation/inhibition
Introduction
The G-LISA series of Small G-Protein Activation Assays are ELISA based assays with which you can measure the GTP form of small G-proteins from lysates of cells or tissues and all in less than 3 h. The level of activation is measured with luminescence. For a more detailed introduction on G-LISA™ assays and a listing of other available G-LISA™ kits, see our main G-LISA™ page. The Rac1 G-LISA™ Activation Assay (Cat.# BK126) measures the level of GTP-loaded Rac1 protein in cell lysates, this is in contrast to Cat. BK125 which does not distinguish between activated Rac1,2 or 3. For a kit to measure RhoA activation please check webpage BK124.
The Rac1 G-LISA Activation Assay is very sensitive and has excellent accuracy for duplicate samples. See G-LISA™ FAQs tab on our G-LISA™ page for more details.
Kit contents
The kit contains sufficient reagents to perform 96 Rac activation assays. Since the Rac-GTP affinity wells are supplied as strips and the strips can be broken into smaller pieces, each kit can be used for anywhere from one to multiple assays. The following components are included in the kit:
- 96 Rac-GTP affinity wells (divisible into 96 individual wells)
- Lysis buffer
- Binding buffer
- Antigen presenting buffer
- Wash buffer
- Antibody dilution buffer
- Anti-Rac1 antibody
- HRP-labeled secondary antibody
- Positive control Rac1 protein
- Protease inhibitor cocktail (Cat. # PIC02)
- Luminescence detection reagents
- Precision Red™ Advanced protein assay reagent (Cat. # ADV02)
- Manual with detailed protocols and extensive troubleshooting guide
Equipment needed
- 96-well plate luminometer
- Multichannel or multidispensing pipettor
- Orbital microplate shaker capable of at least 200 rpm shaking (400 rpm is optimal)
Example results
Serum starved Swiss 3T3 cells were stimulated with the Rac activating compound EGF and Rac activation was measured with the G-LISA method (Fig 1 and 2).
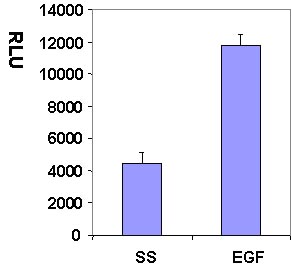
Figure 1. Rac1 activation by EGF measured by G-LISA™ kit BK126. Swiss 3T3 (mouse) cells were serum starved for 24 h and treated with EGF (10ng/ml for 2.5 min) or buffer only (SS). 10 µg of cell lysates were subjected to the G-LISA™ assay. Luminescence measured over 100 milli-second.
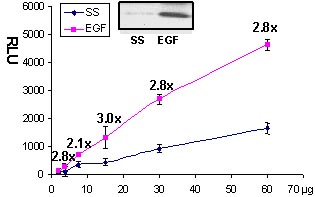
Figure 2. Rac activation by EGF measured by G-LISA™. Swiss 3T3 cells were serum starved (SS) for 24 h and treated with EGF (10 ng/ml for 2 min). 60, 30, 15, 7.5, 2.5 µg of cell lysates were subjected to the G-LISA™ assay. Luminescence was measured over 10 milli-seconds. 500 µg of the same lysates were subjected to the traditional PAK pull-down assay (shown in inset, Cat.# BK035).
G-LISA™ Products:
Cdc42 G-LISA™ Activation Assay, colorimetric format (Cat.# BK127)
Rac1,2,3 G-LISA™ Activation Assay, colorimetric format (Cat.# BK125)
RhoA G-LISA™ Activation Assay, colorimetric format (Cat.# BK124)
RhoA G-LISA™ Activation Assay, luminescence format (Cat.# BK121)
Associated Products:
Anti-Cdc42 monoclonal antibody (Cat.# ACD03)
Anti-Rac1 monoclonal antibody (Cat.# ARC03)
Anti-RhoA monoclonal antibody (Cat.# ARH03)
For product Datasheets and MSDSs please click on the PDF links below. For additional information, click on the FAQs tab above or contact our Technical Support department at tservice@cytoskeleton.com
 G-LISA Activation Assay Technical Guide download here
G-LISA Activation Assay Technical Guide download here For our G-LISA Data Analysis (Luminescence) Excel Template please download here.
For our G-LISA Data Analysis (Luminescence) Excel Template please download here.
Question 1: There is less than a 2-fold difference in signal intensity between my positive control and lysis buffer blank. Why?
Answer 1: To accurately measure luminescence signal intensity between the positive control and buffer blank, please check the instrument settings on the luminometer (see below for suggestions). We also recommend running some “set-up” experiments with just the buffer blank and positive control to determine optimal settings for detecting the positive control signal 3-5 fold higher than the buffer blank. It is also important to remember to use a fresh control protein tube for each run of positive control samples. Do not store and re-use the positive control.
Machine Settings
Gain
Gain controls the sensitivity of the machine. Most luminometers do not allow manual alteration of gain and use an auto-calibration or limited calibration function. Turn off auto-settings and auto-calibration to use the machine in manual mode. It is important to contact the luminometer manufacturer or consult the user’s manual to determine the best way to alter the machine sensitivity. If gain can be altered, one should read at low, medium and high gains to determine the reading within the linear range of the assay (positive control should be 3-5X higher than blank). Gain range varies with instrument. For example, gain in the Tecan GmbH SpectroFluor Plus ranges from 0 - 150 (where 150 is the highest).
Integration Time
This parameter can be varied on most machines. It is a good idea to set the machine at the lowest integration time (usually 10 – 100 ms). Integration times greater than 200 ms are likely to read out of the linear range of the assay and may require lowering of gain or dilution of primary and/or secondary antibodies.
Shaking
Most machines give the shaking option. The recommended setting is 5 sec shake, medium orbital speed before read. This option is not essential to the assay.
Temperature
Room temperature
Plate type
Any setting that specifies 96 well flat, white will be sufficient.
Filters
Luminescence does not require excitation or emission filters so the filter spaces should be left blank. If this is not an option, excitation can be set at any value and emission should be set between 400-500nm, with 430-445 as optimal setting.
Question 2: My arbitrary luminescence units (ALU) or relative luminescence units (RLU) are very different from what is depicted in the manual. Why?
Answer 2: This is very typical as the luminescence units will vary from luminometer to luminometer based on the machine’s sensitivity and instrument settings. The important information to take note of is what the relationship is between buffer blank and positive control luminescence values. The positive control signal should be 3-5 fold greater than the buffer blank luminescence signal. If that is the case, then the G-LISA assay is functioning in the linear range and experimental samples can now be processed.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com.