+3
Loading...
Rabbit skeletal muscle actin AKL99 is labeled with pyrene (N-1-pyrene iodoacetamide) at the reactive cysteine 374 residue:
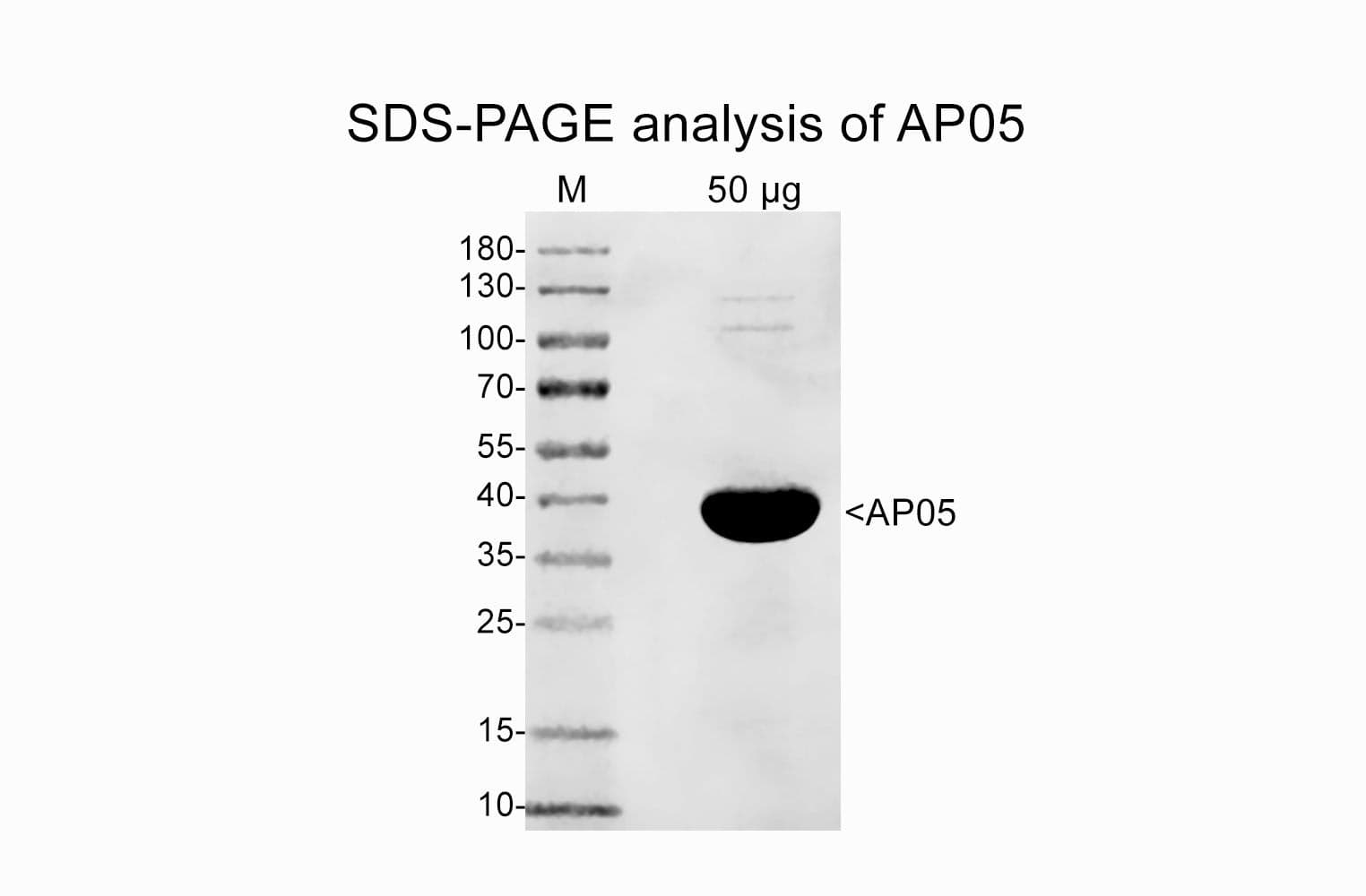
Protein purity is assessed by scanning densitometry of Coomassie Blue stained protein on a12% polyacrylamide gel. Purity was determined to be >99% pure
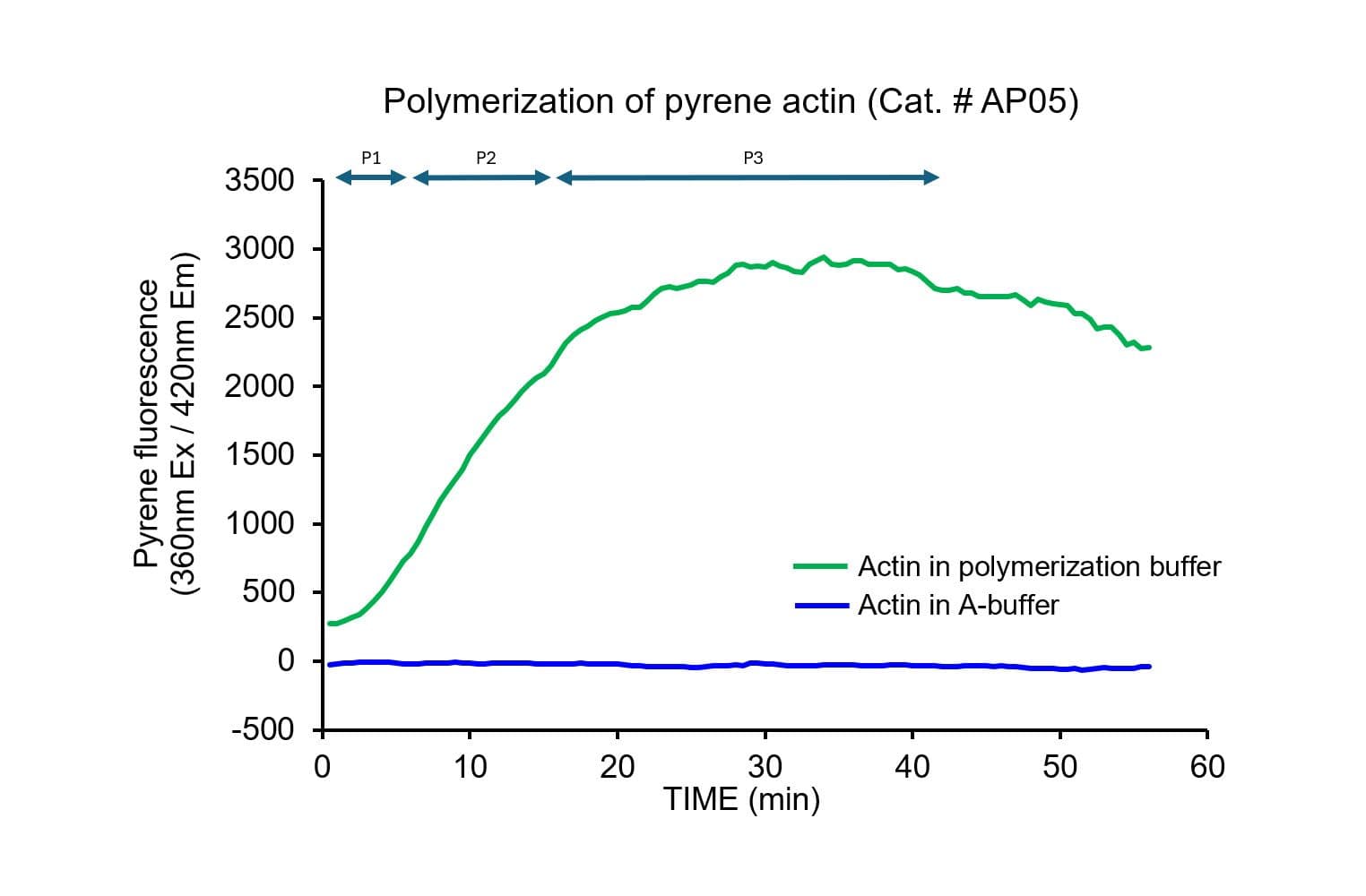
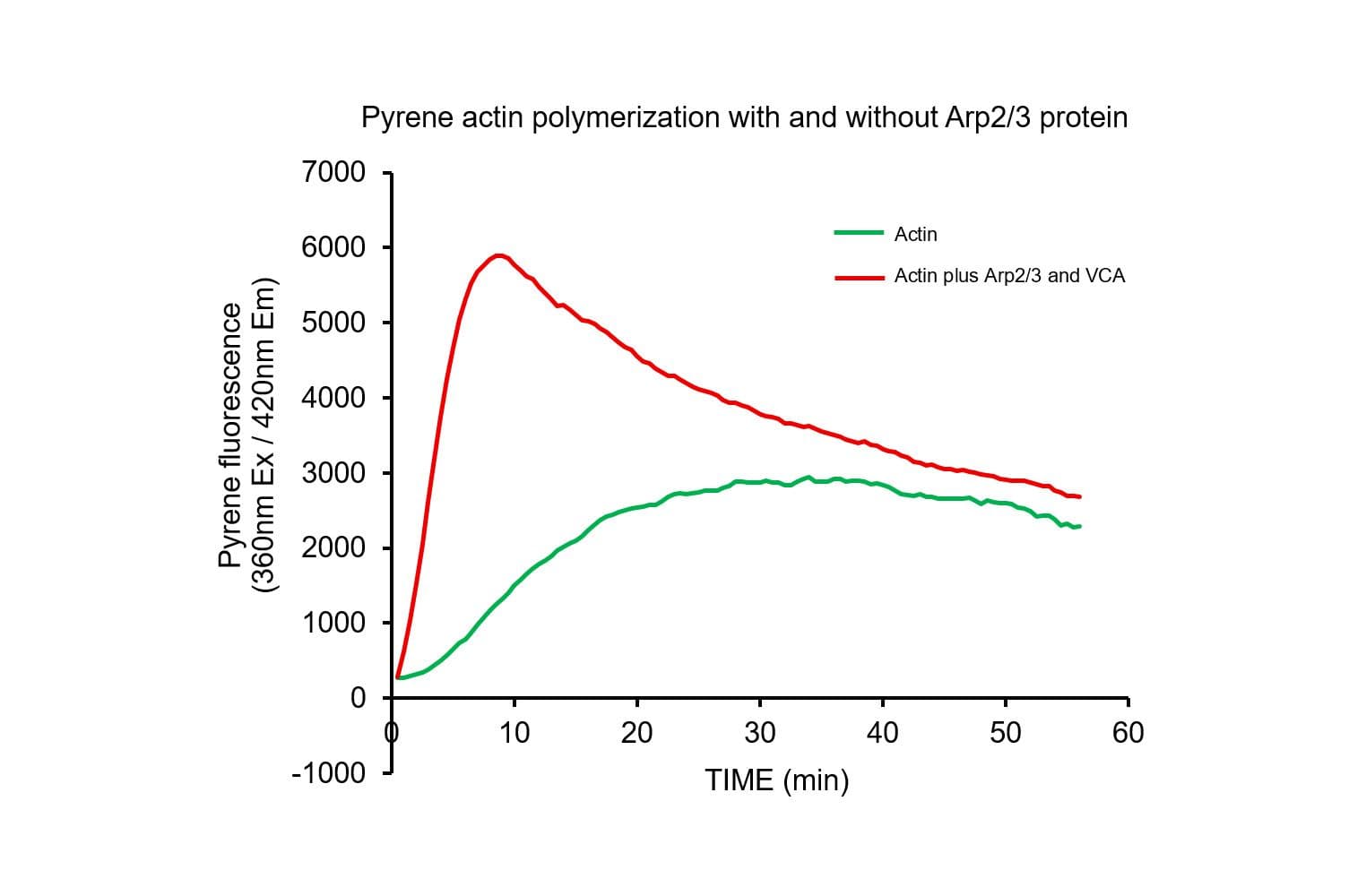
The biological activity of pyrene-labeled actin has been determined by its ability to polymerize into filaments in vitro efficiently. Pyrene actin exhibits a 7–12-fold increase in fluorescence (Ex: 360 nm, Em: 405 nm) upon polymerization, equivalent to >80% polymerization.
Cat. #AP05