Protocol
+3
Loading...
Cat. #ADV02
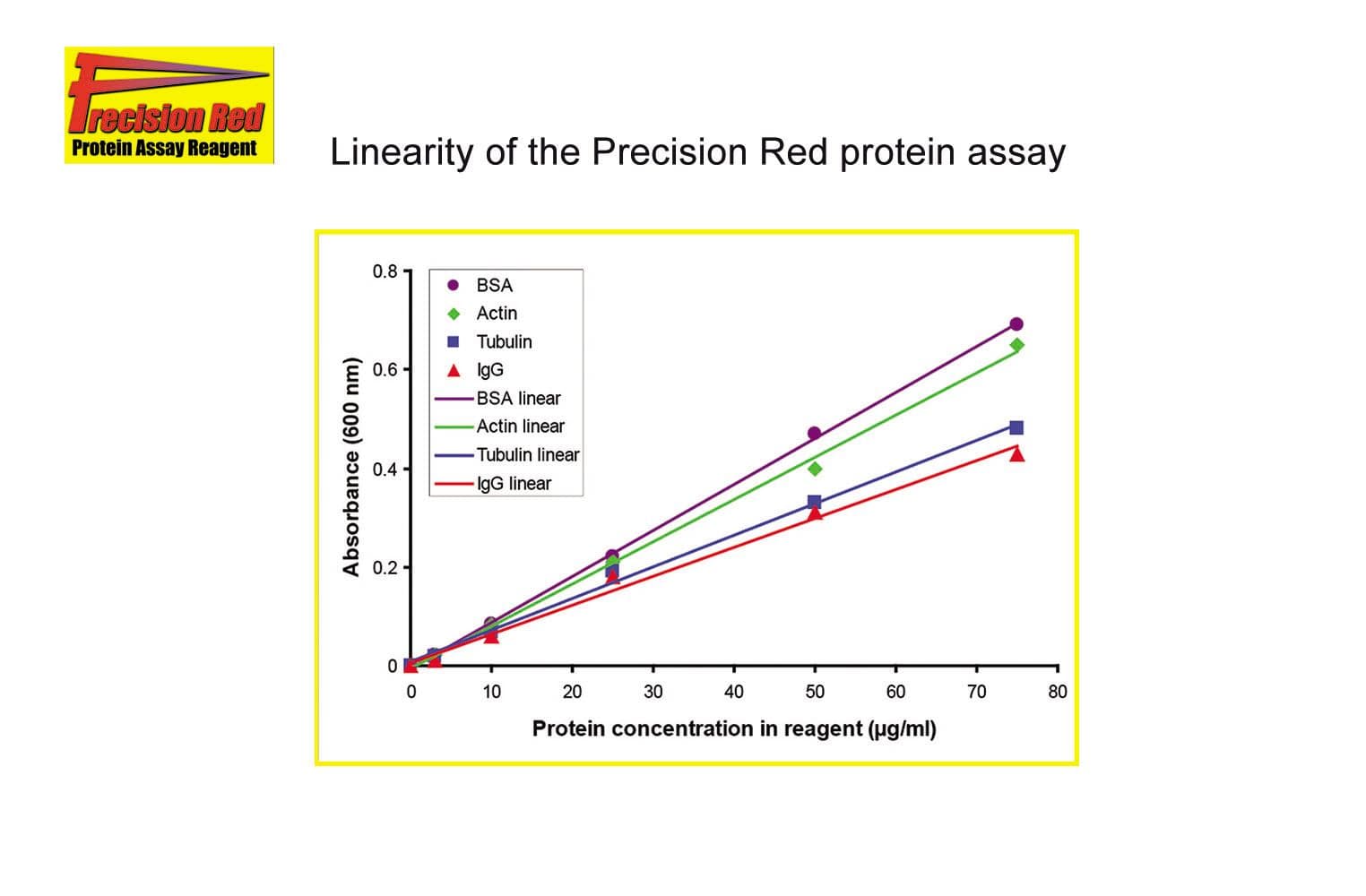

ADV02 provides a rapid, sensitive, and reproducible method for quantifying purified proteins or complex protein mixtures. Designed with a simple, colorimetric readout, this assay delivers accurate results across a broad concentration range, making it ideal for routine protein quantitation in biochemical and cell biology applications. Its streamlined protocol reduces hands-on time while maintaining high precision, ensuring reliable performance.
Key characteristics
Manufactured using only ACS reagents of ≥99% purity
The biological activity of ADV02 is validated by its ability to measure protein concentration in the linear range of 0.25-1.5 mg/ml, cv ≤3%