+3
Loading...
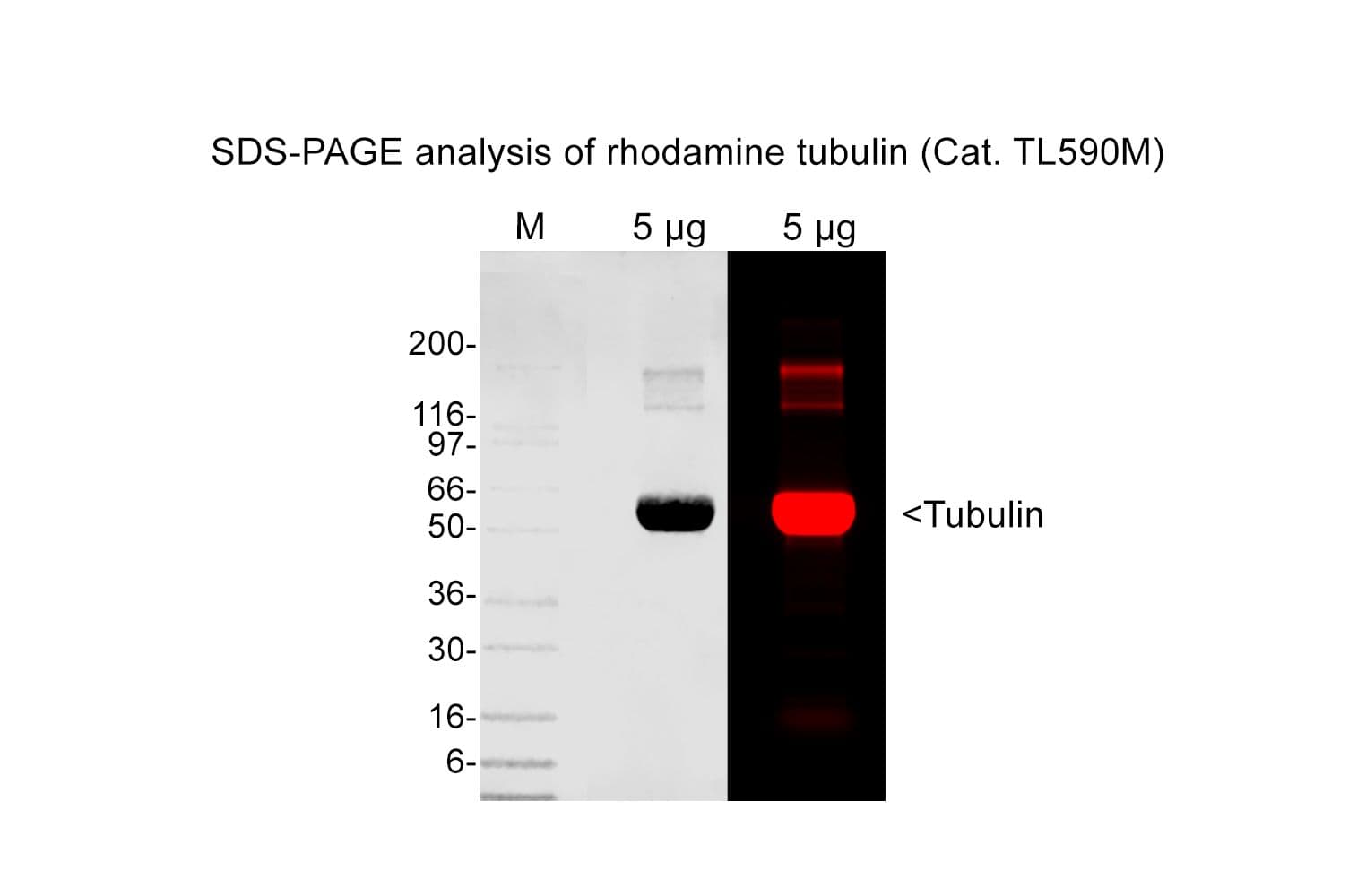
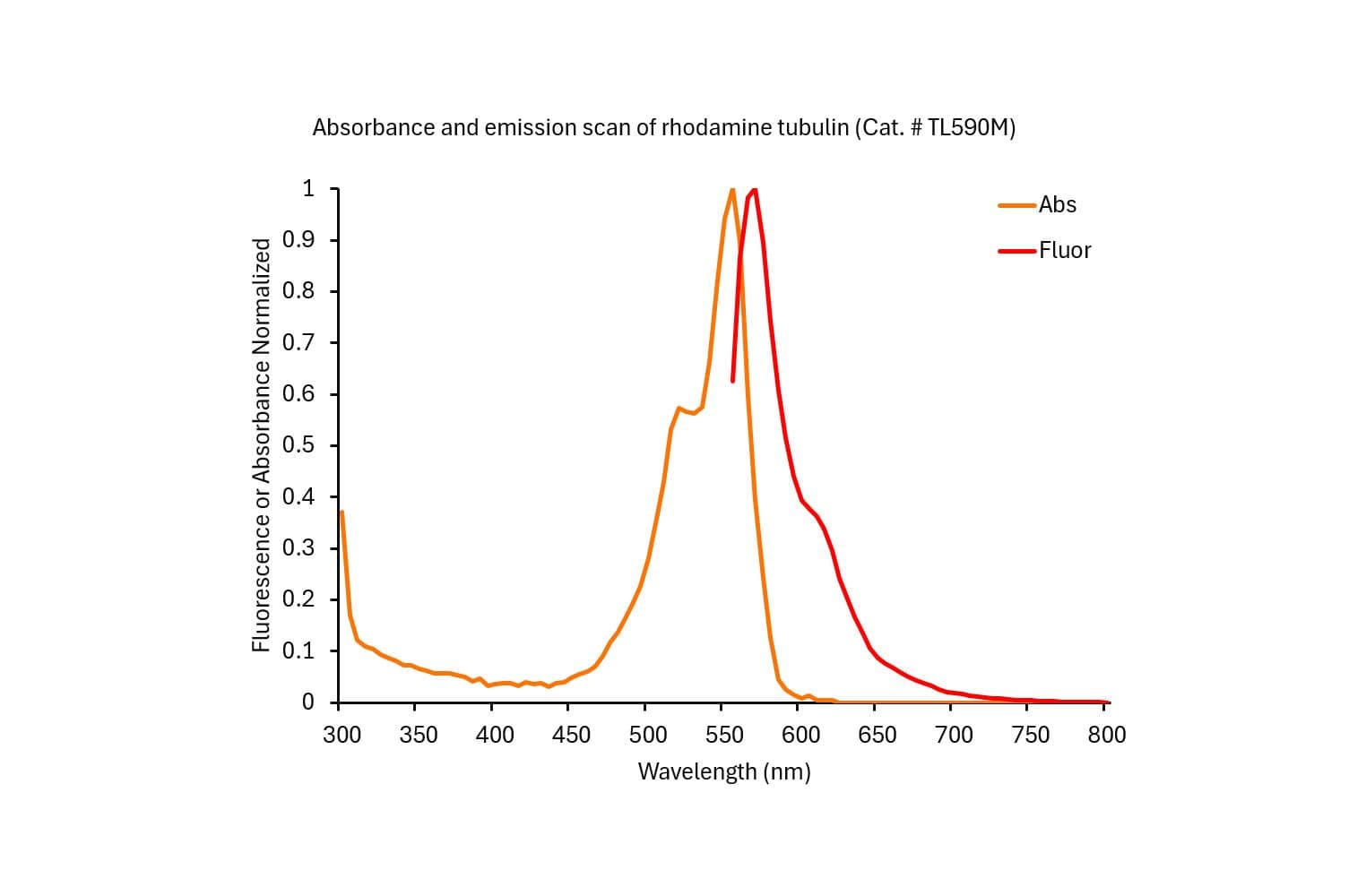
Tubulin protein T240 is labeled on random surface lysines using an activated ester of rhodamine;
Protein purity is assessed by scanning densitometry of Coomassie Blue stained protein on a12% polyacrylamide gel. Purity is determined to be >99% pure.
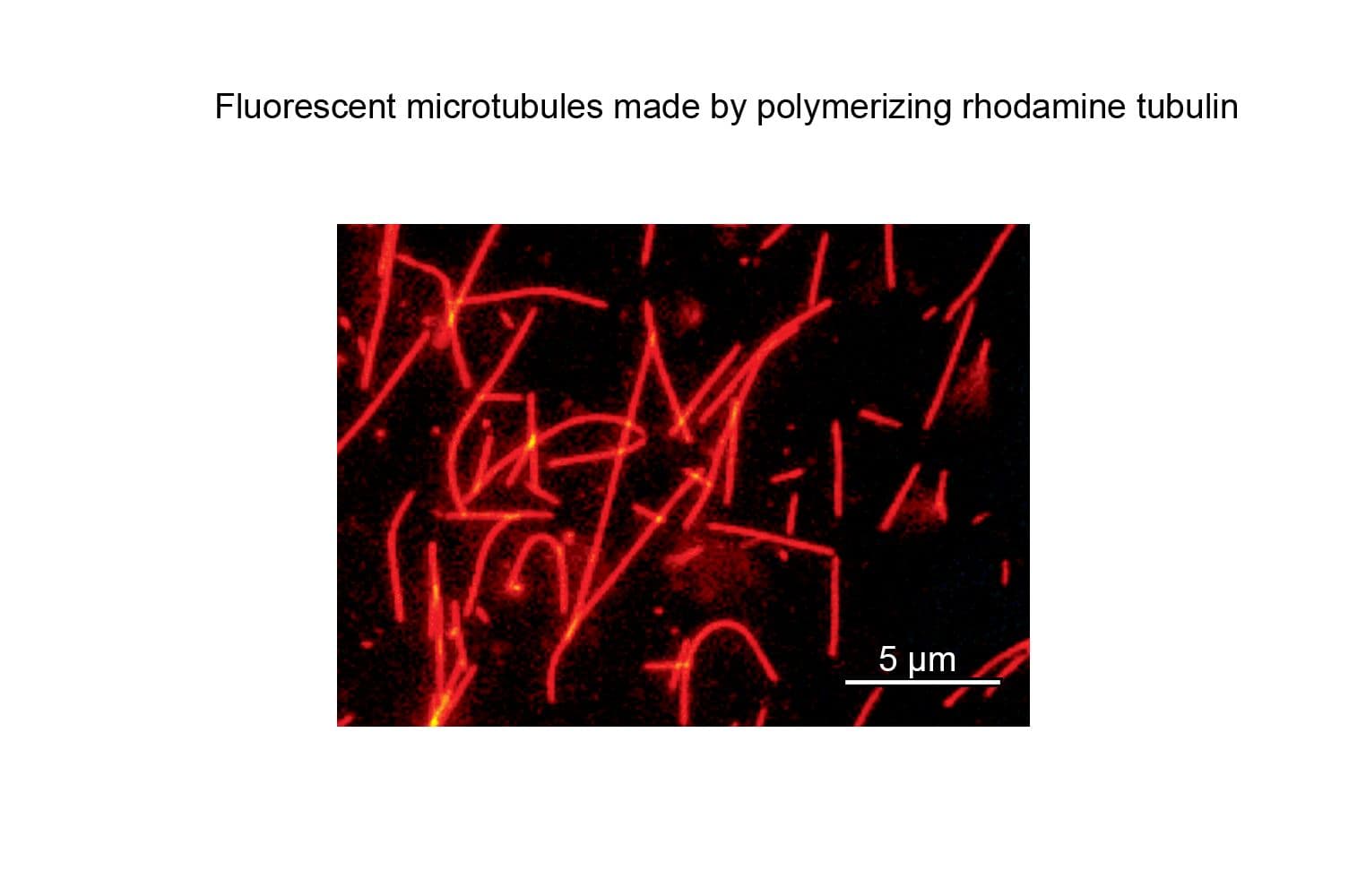
TL590M tubulin retains high biological activity, with >80% polymerizing into microtubules in vitro, as confirmed by a spin-down assay. This performance is comparable to that of unmodified tubulin.
Cat. #TL590M