RasGRF1 GEF Protein (Cdc25 Exchange Domain, aa1038-1270, MBP tag)
Cat. #CS-GE03

Cat. #CS-GE03



RasGRF1 is a guanine nucleotide exchange factor (GEF) that activates Ras proteins, playing a key role in transmitting signals from cell surface receptors to downstream pathways controlling cell growth and differentiation. It is particularly important in neuronal signaling and learning processes and is also implicated in regulating pancreatic β-cell function and insulin secretion.
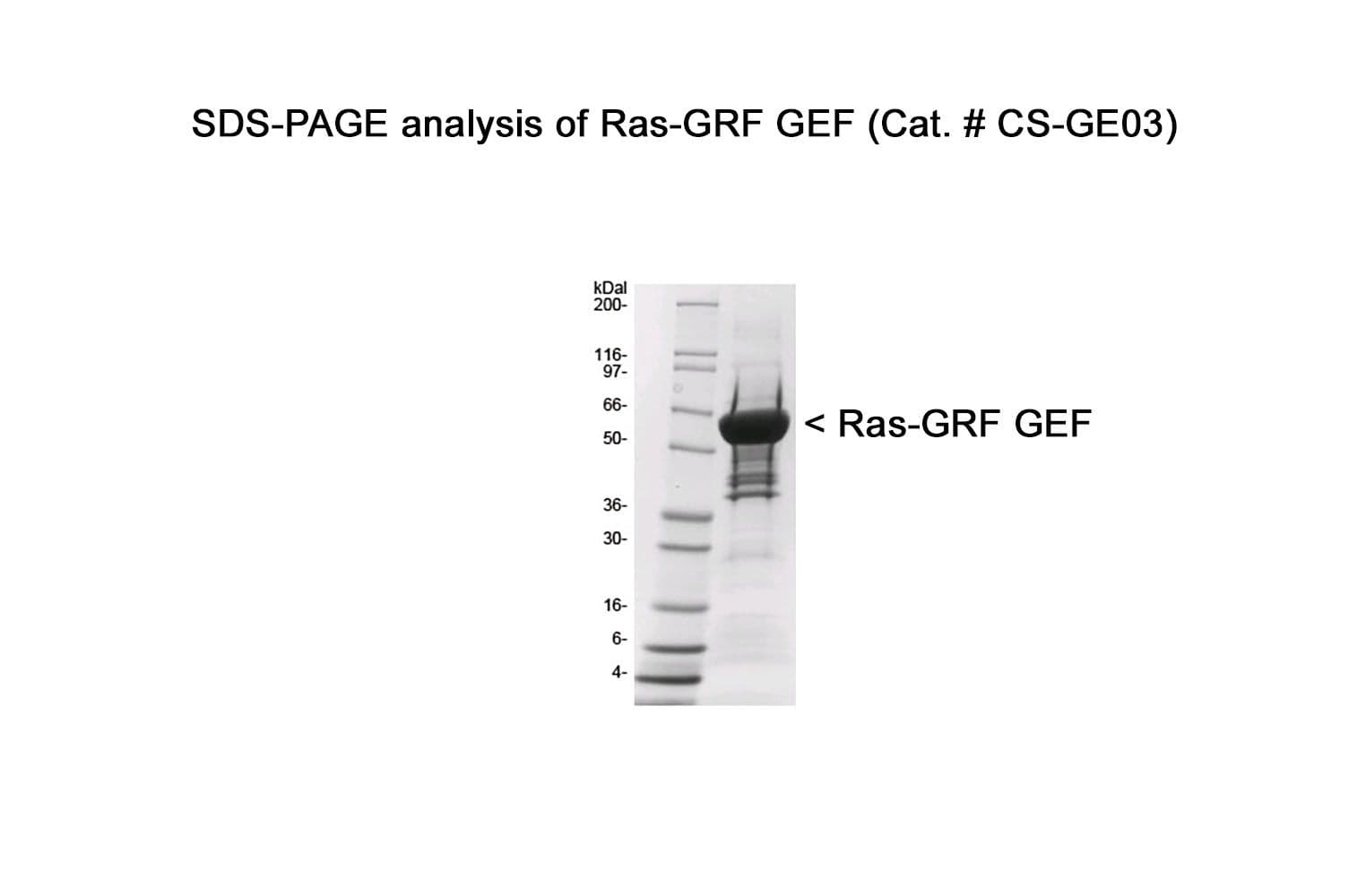
The Cdc25 domain of human RasGRF1 protein (aa 1038-1270) has been produced in a bacterial expression system. It contains a maltose binding protein (MBP) fusion at its amino terminus. The molecular weight of MBP-RasGRF1-Cdc25 binding domain is approximately 60 kDa. The protein is supplied as a lyophilized powder.
Protein purity is assessed using scanning densitometry of Coomassie-stained SDS-PAGE gels. CS-GE03 is ≥85% pure
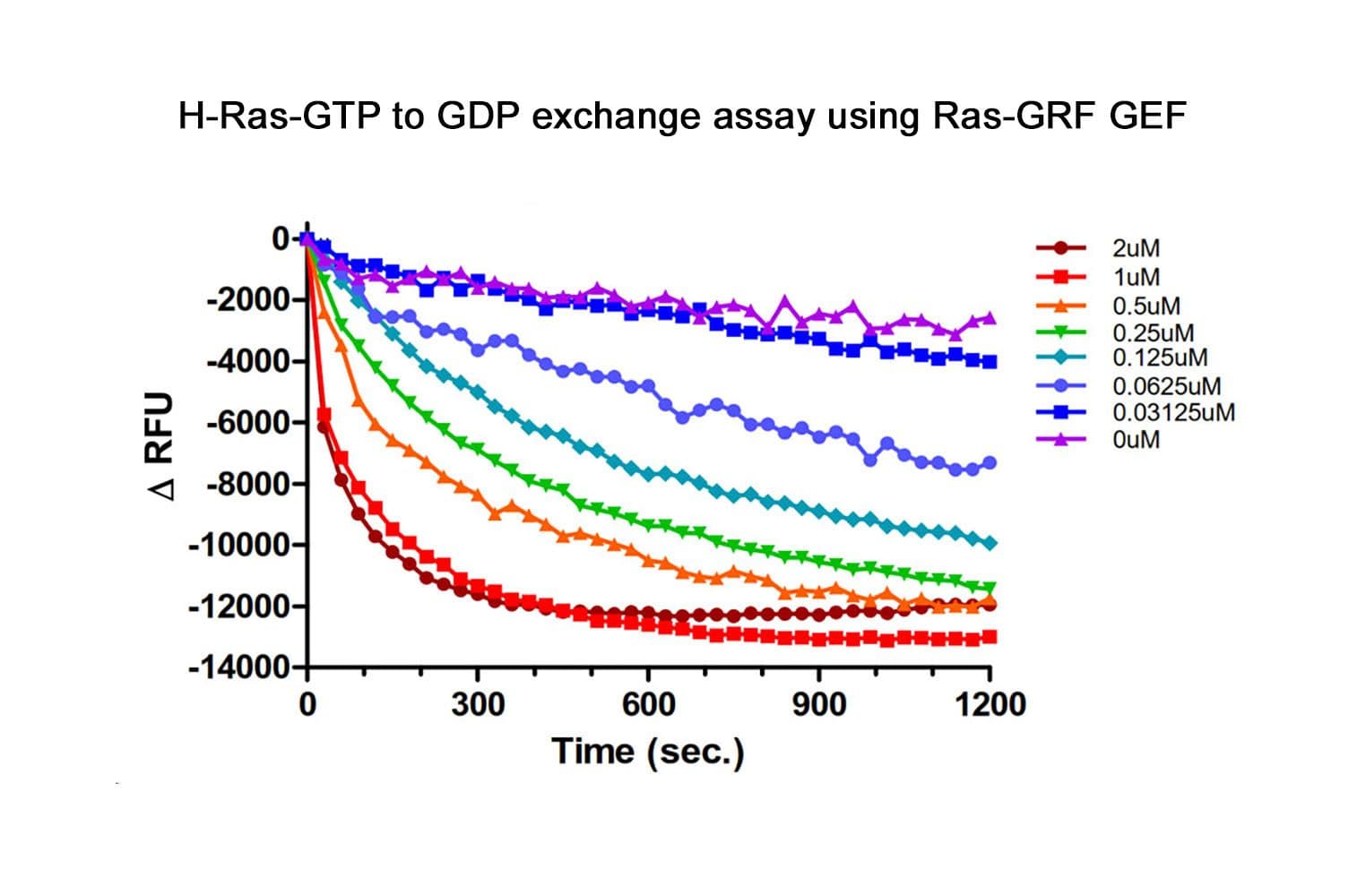
The biological activity of the Cdc25 (GEF) domain of RasGRF1 is determined using an in vitro GEF assay. Under the experimental conditions (see datasheet), CS-GE03 increases H-Ras, N-Ras and K-Ras GTP exchange by ≥5 fold over intrinsic Ras exchange.