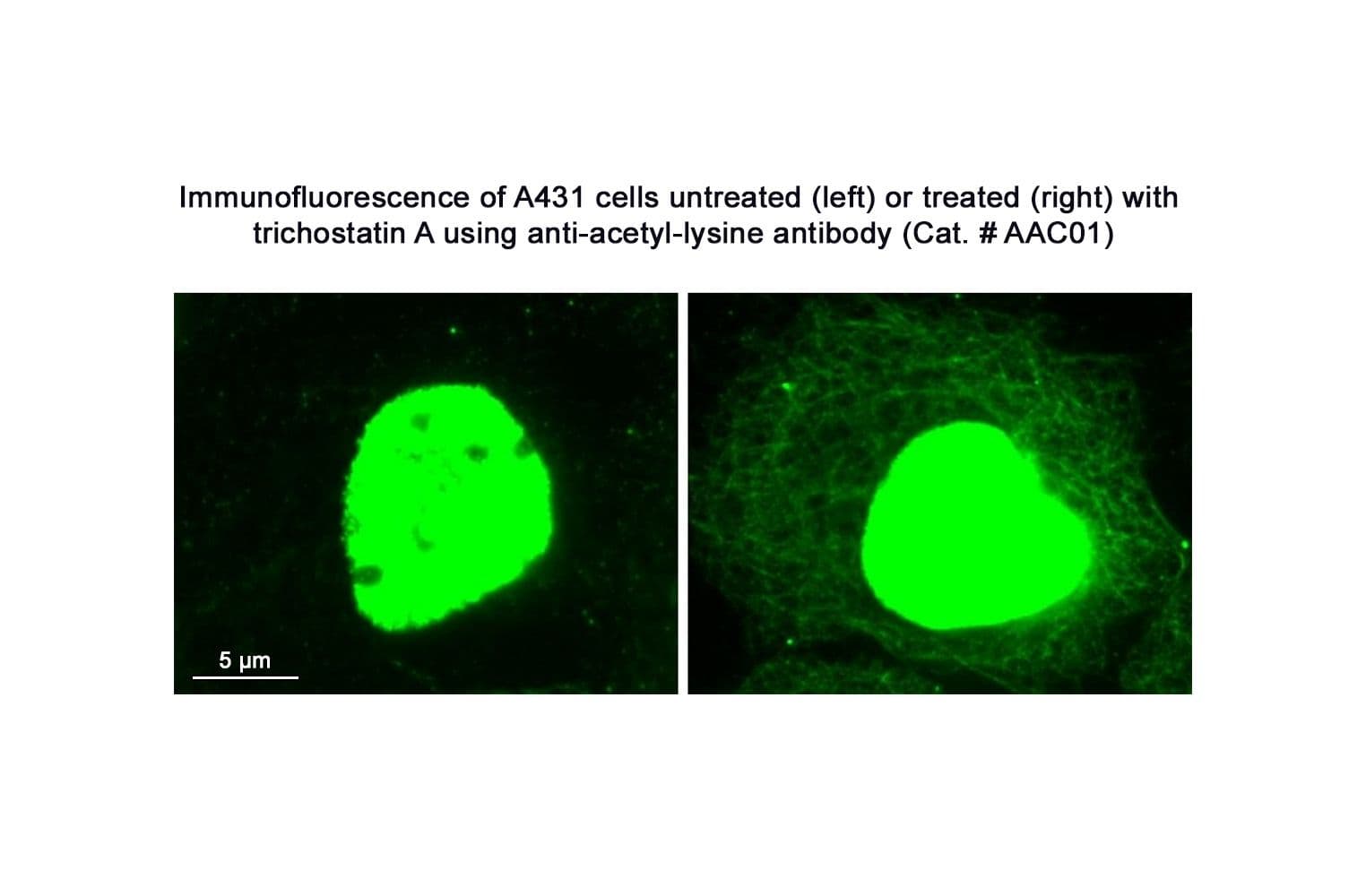
Anti-acetyl lysine antibody (AAC01) was raised against a proprietary mix of acetylated proteins to ensure broad acetyl lysine recognition. It detects a wide range of acetylated proteins, including tubulin, histones, glutamic dehydrogenase, actins, and Arp 2/3 complex proteins. AAC01 is purified via Protein G chromatography.
Host/isotype/clone:
- Mouse/IgG2b
- Clone: 3C6.08.20
Reactivity:
- All species (acetyl-lysine is identical in all species)
Amount of material:
- AAC01: 2 x 100 µl when reconstituted (100-180 µg per tube)
- AAC01-S: 1 X 25 µl when reconstituted (25-45 µg per tube)
Working concentrations:
- WB 1:1000 dilution 1-1.8 µg/ml final 200 ml working volume - full size Ab
- IF 1:200 dilution 5-9 µg/ml final 40 ml working volume - full size Ab
- IP 20 ul neat 20-36 µg per IP 10 reactions – full size Ab
- ChIP 1:100 dilution 10-18 µg/ml final 20 ml working volume – full size Ab
Formulation:
- Supplied as a lyophilized powder
- Buffer composition after reconstitution in 50% glycerol in water: 250 mM PIPES pH 7.0, 1% sucrose, 0.5% dextran, 50% glycerol.