Protocol
+3
Loading...
Cat. #AUB01
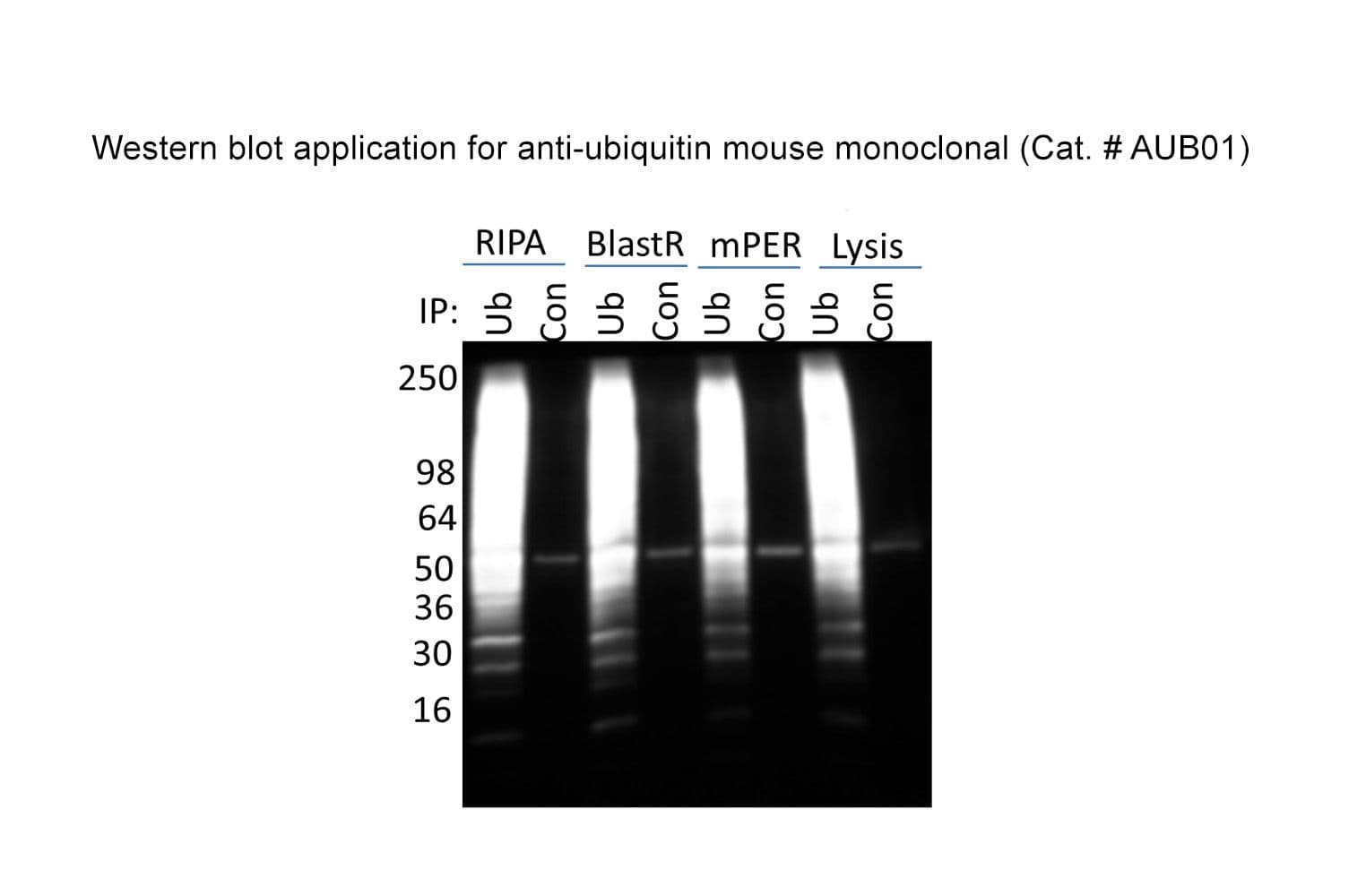
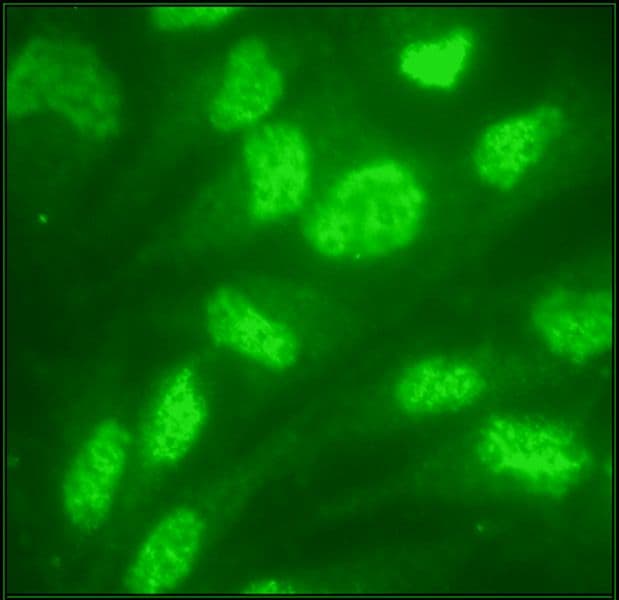

Anti-ubiquitin antibody (AUB01) was raised against full-length bovine ubiquitin protein. It detects a broad profile of ubiquitinated proteins and free ubiquitin in cell lysates. AUB01 is purified by Protein G affinity chromatography. The clone is licensed from Fred Hutchinson Cancer Research Center; the antibody is made in-house by Cytoskeleton.
Host/isotype/clone:
Reactivity:
Amount of material:
Working concentrations:
Formulation
Sensitivity:
Specificity: