Introduction
Microtubules are hollow cylindrical structures found in all eukaryotic cells,1 displaying a wide range of structural and functional specialization. They are dynamic polymers built from dimeric building blocks consisting of one α-tubulin and one β-tubulin subunit. Humans have nine α-tubulin and nine β-tubulin genes,2 and specific expression patterns of these tubulin isotypes underlie the varied structural and mechanical properties of microtubules found in different cell types.3
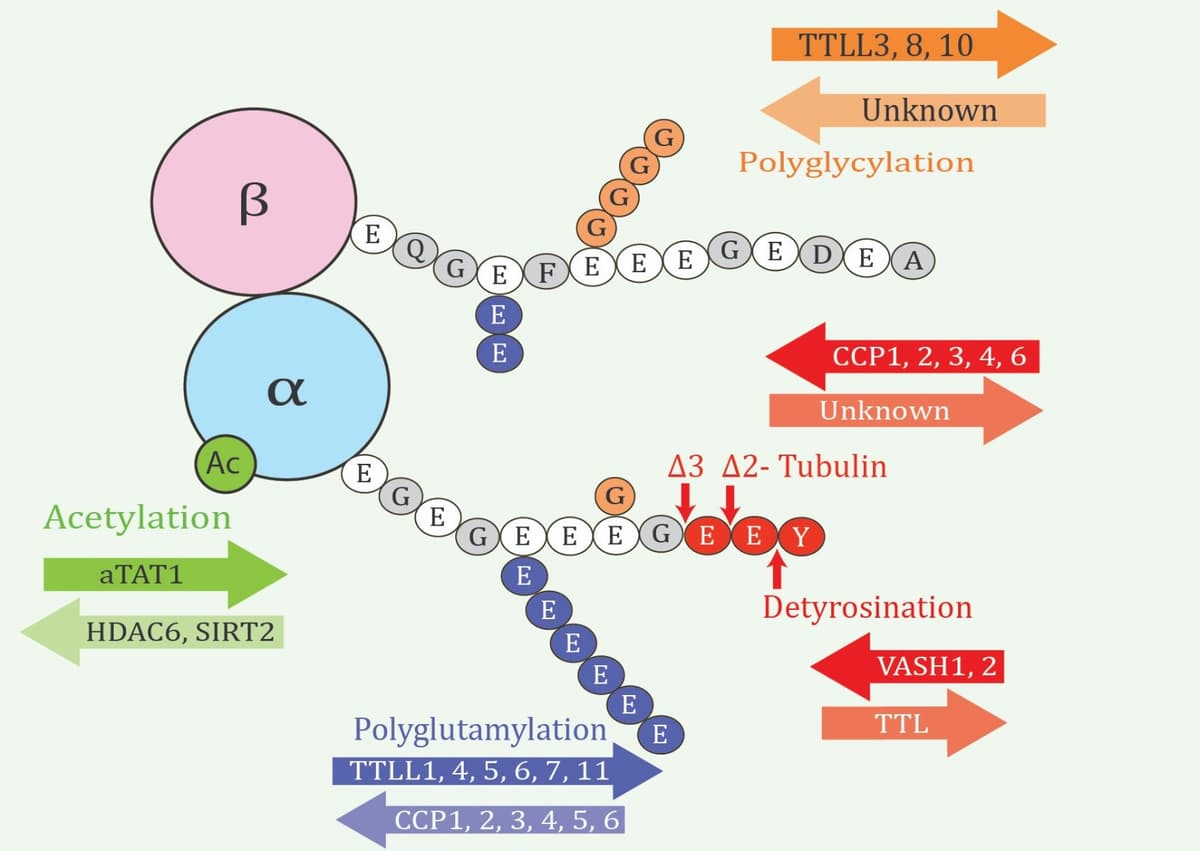
Tubulin genetic diversity is complemented by a wide range of posttranslational modifications (PTMs) (Figure 1 ) that also vary considerably between cell types, as well as exhibiting spatial and developmental regulation.4 This multiplicity of tubulin isotypes and modifications led to the concept of the “tubulin code,” analogous to the histone code, which encompasses “the diversity of tubulin isoforms and PTMs that can combine in numerous ways to confer unique functionality to subsets of microtubules within the cell”.5
In this newsletter, we will review some prominent tubulin PTMs alongside more recent discoveries such as methylation, whose role in normal and pathological processes is still largely unknown. We will also see how advances in imaging techniques are enabling mapping of tubulin PTMs at unprecedented resolution, uncovering fascinating new insights into microtubule specialization and function.

α-Tubulin Acetylation
While multiple tubulin acetylation sites have been identified, the best-studied example is the highly conserved Lys40 of α-tubulins.6,7 Noted for its unique location in the microtubule inner lumen,2 this prominent PTM is written by α-tubulin N-acetyltransferase 1 (ATAT1) and erased by HDAC6 or SIRT2.
Historically, Lys40 acetylation was considered a stabilizing modification and a marker of long-lived microtubules, largely due to the high levels observed in neurons.8 Recent data inform a more nuanced understanding where it enhances flexibility of microtubules by modifying interactions between protofilaments, protecting against mechanical aging and breakage with repeated bending.6 This explains the abundant Lys40 acetylation found on ciliary axonemes.9
Conversely, tubulin deacetylation (mediated by HDAC6) is important during platelet activation.10 The marginal band, a coiled microtubule structure that gives platelets their characteristic discoid shape, is acetylated at Lys40 in resting platelets but rapidly deacetylated upon platelet activation.11 Tubulin hyperacetylation modifies cytoskeletal morphology during activation and inhibits platelet aggregation; thus, altered microtubule acetylation may be a pathogenic feature of bleeding disorders.12
Another acetylated α-tubulin residue, Lys394, was more recently shown to be essential for microtubule stability in neurons,6 playing a role in correct synaptic morphogenesis.13 Interestingly, this site is not a substrate for ATAT1 and the responsible acetyltransferase is presently unknown.
Tubulin Methylation
In recent years, several histone methyltransferases have also been found to act on tubulin. The first was SETD2, which methylates α-tubulin on Lys40 to write the αTubK40me3 mark that peaks during mitosis and cytokinesis.14 Since haploinsufficiency of SETD2 has been linked to autism spectrum disorder, a recent mouse study examined heterozygous animals bearing a mutation that abolishes tubulin (but not histone) methylation, observing an anxiety-like phenotype in mutant mice.15 Thus, tubulin methylation defects may be the underlying mechanism of certain pathogenic SETD2 mutations. The αTubK40me3 modification was observed on axonal microtubules and appeared to be functionally important for dendritic branching.
Another methyltransferase, SMYD3, was recently found to also methylate Lys40,16 although its functional relationship with SETD2 is not yet clear. The related protein SMYD2 methylates tubulin at a different site,16 αTubK394me3 (intriguingly, also a known acetylation site), and this trimethylated form localizes to the basal body where it regulates ciliogenesis, or more specifically, ciliary length.
α-Tubulin is also monomethylated at Lys311 by KMT5A/SET8,18 an enzyme otherwise associated with the mitotic H4K20me1 histone mark. While KMT5A is implicated in cell cycle regulation and is most highly expressed during mitosis, the role of αTubK311me1 in these processes is as yet undetermined. Multiple other methylation sites have been identified and await functional investigation, including α-tubulin Arg2, Arg79, Lys124, Lys166, and Lys 304, along with β tubulin Arg2, Lys19, Arg86, Lys297, and Arg318.18,19
Importantly, modification by SETD2 (and presumably other methyltransferases) occurs preferentially on soluble tubulin, not on assembled microtubules as for ATAT1.20 This explains how regions of methylated tubulin can be spatially distinct from those of acetylated tubulin,21 suggesting coordinated formation of functionally differentiated microtubule arrays in the cell.
The Tyrosination/Detyrosination Cycle
Most α-tubulin isotypes carry a C-terminal Tyr residue that may be enzymatically removed or religated in a circular catalytic cycle. Tyrosination and detyrosination are associated with dynamic and stable microtubules, respectively, although these characteristics reflect differential interactions with microtubule-associated proteins (MAPs) rather than changes in inherent microtubule dynamics.22 Many motor proteins are sensitive to tyrosination status,2 including the kinetochore motor CENP-E that guides chromosome alignment during mitosis in a detyrosination-dependent manner.
The Tyr religation step is performed solely by tubulin tyrosine ligase (TTL), discovered in 1975, but the enzymes responsible for detyrosination were not identified until relatively recently.2 The vasohibins, VASH1 and VASH2, processively detyrosinate α-tubulin in intact microtubules with different kinetics to generate distinct modification patterns.23 Specifically, VASH1 shows faster turnover and leads to diffuse detyrosinated areas, whereas VASH2 exhibits much longer residence times, resulting in more localized detyrosination.
Aberrant detyrosination has been implicated in the pathology of heart failure.5 In one report, the microtubule network was examined in cardiomyocytes from failing hearts (after transplant) and non-failing hearts (from cadaveric donors), revealing a significant increase in microtubule density and detyrosination associated with the disease.24 These findings suggested impaired microtubule dynamics in heart failure, underlying increased myocyte stiffness and impaired contractility.5
During discovery of the vasohibins,25 experimental evidence pointed to at least one “missing” detyrosinase that was later identified as MATCAP1,26 a previously uncharacterized metalloprotease with subtly different substrate preferences to VASH2. This enzyme can also directly generate Δ2-tubulin, an irreversible α-tubulin PTM normally formed through downstream processing of detyrosinated tubulin by cytosolic carboxypeptidases (CCPs). While the function of the Δ2 form is not well understood, it is chiefly associated with hyperstable microtubules in neurons.27
Precise Spatial Patterning of Microtubule PTMs
Advances in super-resolution imaging are allowing mapping of multiple PTMs at the single-cell level, or even on individual microtubules.28,29 Two such studies found distinct radially distributed microtubule populations in neuronal dendrites: one highly acetylated and mostly detyrosinated in the center, and the other mostly tyrosinated with little acetylation at the periphery, near the plasma membrane.30,31
The tyrosinated outer region may serve as a specialized responsive scaffold facilitating branch dynamics and synaptic plasticity,2 for which tyrosinated tubulin in dendritic spines is especially important.32 Given that decreased TTL expression is seen in Alzheimer’s disease, an imbalanced tyrosination/detyrosination cycle may contribute to synaptic dysfunction through perturbed microtubule dynamics.
Another recent example highlights precise nanopatterning of PTMs on axonemal microtubule doublets in motile cilia, where the B-tubules are extensively glycated except on one specific protofilament, which is polyglutamylated instead.29 This arrangement is conserved between Chlamydomonas reinhardtii and mouse and is essential for ciliary beating.33 By restricting localization of the nexin–dynein regulatory complex (NDRC) to the polyglutamylated protofilament through electrostatic interactions, a sliding joint is created that mechanically stabilizes the axoneme during bending.29
Understudied Tubulin Modifications
Surprisingly, tubulin phosphorylation has received relatively little attention despite the many target sites identified in proteomics studies.34 The best-studied case is Ser172 in β-tubulin, which is phosphorylated by either CDK1 or DYRK1A.2 Mutations of this residue in TUBB2B (β2b-tubulin) disrupt cell migration and axon outgrowth, causing structural brain abnormalities in bilateral asymmetrical polymicrogyria;35 cellular studies in yeast confirmed the importance of Ser172 for normal functioning of cytoplasmic and spindle microtubules.36
Several other tubulin PTMs await detailed characterization, including SUMOylation on α1-, α6-, and β1-tubulins,34 which promotes microtubule disassembly, and mono-ADP-ribosylation (MARylation) of α-tubulin, recently identified in cancer cells and associated with increased migration.37
Conclusions and Future Outlook
The above sections have provided a brief snapshot of the emerging complexity of the tubulin code. While some individual PTMs are well understood and others remain relatively unexplored, the key challenge moving forward will be to interpret the multiplicity of PTMs that can occur even on the same microtubule. These combinations are presumed to be functionally relevant, albeit much less studied than the individual modifications.
For example, acetylation, detyrosination, and Δ2-tubulin are all present in microtubules of the platelet marginal band.12 In neurons, methylation tends to colocalize with tyrosination and acetylation coincides with detyrosination, and crosstalk between these PTMs is seemingly disrupted in Alzheimer’s disease.7,15 The most recently discovered tubulin detyrosinase, MATCAP1, also acts as a deglutamylase, adding a further layer of complexity to regulation.26 Future studies should continue to elucidate the intricate cellular language of the tubulin code, particularly addressing the origins of precise spatial regulation and nanopatterning of PTMs, which currently remains unexplained.29
References
1. Sawin KE, Tran PT. Cytoplasmic microtubule organization in fission yeast. Yeast. 2006;23(13):1001–1014. https://doi.org/10.1002/yea.1404.
2. McKenna ED, Sarbanes SL, Cummings SW, Roll-Mecak A. The tubulin code, from molecules to health and disease. Annu Rev Cell Dev Biol. 2023;39:331–361. https://doi.org/10.1146/annurev-cellbio-030123-032748.
3. Gasic I. Regulation of tubulin gene expression: from isotype identity to functional specialization. Front Cell Dev Biol. 2022;10:898076. https://doi.org/10.3389/fcell.2022.898076.
4. Roll-Mecak A. The tubulin code in microtubule dynamics and information encoding. Dev Cell. 2020;54(1):7–20. https://doi.org/10.1016/j.devcel.2020.06.008.
5. Caporizzo MA, Chen CY, Prosser BL. Cardiac microtubules in health and heart disease. Exp Biol Med (Maywood). 2019;244(15):1255–1272. https://doi.org/10.1177/1535370219868960.
6. Carmona B, Marinho HS, Matos CL, Nolasco S, Soares H. Tubulin post-translational modifications: the elusive roles of acetylation. Biology (Basel). 2023;12(4):561. https://doi.org/10.3390/biology12040561.
7. Martínez-Hernández J, Parato J, Sharma A, et al. Crosstalk between acetylation and the tyrosination/detyrosination cycle of α-tubulin in Alzheimer’s disease. Front Cell Dev Biol. 2022;10:926914. https://doi.org/10.3389/fcell.2022.926914.
8. Janke C, Magiera MM. The tubulin code and its role in controlling microtubule properties and functions. Nat Rev Mol Cell Biol. 2020;21(6):307–326. https://doi.org/10.1038/s41580-020-0214-3.
9. Mahecic D, Gambarotto D, Douglass KM, et al. Homogeneous multifocal excitation for high-throughput super-resolution imaging. Nat Methods. 2020;17(7):726–733. https://doi.org/10.1038/s41592-020-0859-z.
10. Sadoul K, Wang J, Diagouraga B, et al. HDAC6 controls the kinetics of platelet activation. Blood. 2012;120(20):4215–4218. https://doi.org/10.1182/blood-2012-05-428011.
11. Aslan JE, Phillips KG, Healy LD, Itakura A, Pang J, McCarty OJT. Histone deacetylase 6-mediated deacetylation of α-tubulin coordinates cytoskeletal and signaling events during platelet activation. Am J Physiol Cell Physiol. 2013;305(12):C1230–C1239. https://doi.org/10.1152/ajpcell.00053.2013.
12. Magiera MM, Singh P, Gadadhar S, Janke C. Tubulin posttranslational modifications and emerging links to human disease. Cell. 2018;173(6):1323–1327. https://doi.org/10.1016/j.cell.2018.05.018.
13. Saunders HAJ, Johnson-Schlitz DM, Jenkins BV, Volkert PJ, Yang SZ, Wildonger J. Acetylated α tubulin K394 regulates microtubule stability to shape the growth of axon terminals. Curr Biol. 2022;32(3):614–630. https://doi.org/10.1016/j.cub.2021.12.012.
14. Park IY, Powell RT, Tripathi DN, et al. Dual chromatin and cytoskeletal remodeling by SETD2. Cell. 2016;166(4):950–962. https://doi.org/10.1016/j.cell.2016.07.005.
15. Koenning M, Wang X, Karki M, et al. Neuronal SETD2 activity links microtubule methylation to an anxiety-like phenotype in mice. Brain. 2021;144(8):2527–2540. https://doi.org/10.1093/brain/awab200.
16. Agborbesong E, Zhou JX, Zhang H, Li LX, Harris PC, Calvet JP, Li X. Overexpression of SMYD3 promotes autosomal dominant polycystic kidney disease by mediating cell proliferation and genome instability. Biomedicines. 2024;12(3):603. https://doi.org/10.3390/biomedicines12030603.
17. Li LX, Zhou JX, Wang X, Zhang H, Harris PC, Calvet JP, Li X. Cross-talk between CDK4/6 and SMYD2 regulates gene transcription, tubulin methylation, and ciliogenesis. Sci Adv. 2020;6(44):eabb3154. https://doi.org/10.1126/sciadv.abb3154.
18. Chin HG, Esteve P-O, Ruse C, Lee J, Schaus SE, Pradhan S, Hansen U. The microtubule-associated histone methyltransferase SET8, facilitated by transcription factor LSF, methylates α-tubulin. J Biol Chem. 2020;295(14):4748–4759. https://doi.org/10.1074/jbc.ra119.010951.
19. Ceglowski J, Hoffman HK, Neumann AJ, Hoff KJ, McCurdy BL, Moore JK, Prekeris R. TTLL12 is required for primary ciliary axoneme formation in polarized epithelial cells. EMBO Rep. 2024;25(1):198–227. https://doi.org/10.1038/s44319-023-00005-5.
20. Kearns S, Mason FM, Rathmell WK, Park IY, Walker C, Verhey KJ, Cianfrocco MA. Molecular determinants for α-tubulin methylation by SETD2. J Biol Chem. 2021;297(1):100898. https://doi.org/10.1016/j.jbc.2021.100898.
21. MacTaggart B, Kashina A. Posttranslational modifications of the cytoskeleton. Cytoskeleton (Hoboken). 2021;78(4):142–173. https://doi.org/10.1002/cm.21679.
22. Chen J, Kholina E, Szyk A, Fedorov VA, Kovalenko I, Gudimchuk N, Roll-Mecak A. α-Tubulin tail modifications regulate microtubule stability through selective effector recruitment, not changes in intrinsic polymer dynamics. Dev Cell. 2021;56(14):2016–2028. https://doi.org/10.1016/j.devcel.2021.05.005.
23. Ramirez-Rios S, Choi SR, Sanyal C, et al. VASH1–SVBP and VASH2–SVBP generate different detyrosination profiles on microtubules. J Cell Biol. 2023;222(2):e202205096. https://doi.org/10.1083/jcb.202205096.
24. Chen CY, Caporizzo M, Bedi K, et al. Suppression of detyrosinated microtubules improves cardiomyocyte function in human heart failure. Nat Med. 2018;24(8):1225–1233. https://doi.org/10.1038/s41591-018-0046-2.
25. Nieuwenhuis J, Adamopoulos A, Bleijerveld OB, et al. Vasohibins encode tubulin detyrosinating activity. Science. 2017;358(6369):1453–1456. https://doi.org/10.1126/science.aao5676.
26. Nicot S, Gillard G, Impheng H, et al. A family of carboxypeptidases catalyzing α- and β-tubulin tail processing and deglutamylation. Sci Adv. 2023;9(37):eadi7838. https://doi.org/10.1126/sciadv.adi7838.
27. Pero ME, Meregalli C, Qu X, et al. Pathogenic role of delta 2 tubulin in bortezomib-induced peripheral neuropathy. Proc Natl Acad Sci U S A. 2021;118(4):e2012685118. https://doi.org/10.1073/pnas.2012685118.
28. Iwanski MK, Kapitein LC. Cellular cartography: towards an atlas of the neuronal microtubule cytoskeleton. Front Cell Dev Biol. 2023;11:1052245. https://doi.org/10.3389/fcell.2023.1052245.
29. Viar GA, Pigino G. Tubulin posttranslational modifications through the lens of new technologies. Curr Opin Cell Biol. 2024;88:102362. https://doi.org/10.1016/j.ceb.2024.102362.
30. Tas RP, Chazeau A, Cloin BMC, Lambers MLA, Hoogenraad CC, Kapitein LC. Differentiation between oppositely oriented microtubules controls polarized neuronal transport. Neuron. 2017;96(6):1264–1271. https://doi.org/10.1016/j.neuron.2017.11.018.
31. Katrukha EA, Jurriens D, Pastene DMS, Kapitein LC. Quantitative mapping of dense microtubule arrays in mammalian neurons. Elife. 2021;10:e67925. https://doi.org/10.7554/eLife.67925.
32. Peris L, Parato J, Qu X, et al. Tubulin tyrosination regulates synaptic function and is disrupted in Alzheimer’s disease. Brain. 2022;145(7):2486–2506. https://doi.org/10.1093/brain/awab436.
33. Viar GA, Klena N, Martino F, Nievergelt AP, Bolognini D, Capasso P, Pigino G. Protofilament-specific nanopatterns of tubulin post-translational modifications regulate the mechanics of ciliary beating. Curr Biol. 2024;34(19):4464–4475. https://doi.org/10.1016/j.cub.2024.08.021.
34. Peng N, Nakamura F. Microtubule-associated proteins and enzymes modifying tubulin. Cytoskeleton (Hoboken). 2023;80(3–4):60–76. https://doi.org/10.1002/cm.21748.
35. Jaglin XH, Poirier K, Saillour Y, et al. Mutations in the β-tubulin gene TUBB2B result in asymmetrical polymicrogyria. Nat Genet. 2009;41(6):746–752. https://doi.org/10.1038/ng.380.
36. Caudron F, Denarier E, Thibout-Quintana J-C, Brocard J, Andrieux A, Fourest-Lieuvin A. Mutation of Ser172 in yeast β-tubulin induces defects in microtubule dynamics and cell division. PLoS One. 2010;5(10):e13553. https://doi.org/10.1371/journal.pone.0013553.
37. Morone B, Grimaldi G. PARP enzymes and mono-ADP-ribosylation: advancing the connection from interferon-signalling to cancer biology. Expert Rev Mol Med. 2024;26:e17. https://doi.org/10.1017/erm.2024.13.