Lysine acetylation is a post-translational modification (PTM) crucial for regulating the function and localization of many eukaryotic proteins. This PTM is reversible, regulated by histone deacetylases (HDACs) and histone acetyltransferases (HATs). The first evidence of lysine acetylation was discovered in calf thymus histones in 19681. In the following decades, non-histone proteins such as high mobility group protein 1, alpha-tubulin, and tumor suppressor P53 were shown to be lysine acetylated2-4. With the development of pan-acetyl-lysine antibodies and their use in mass spectrometry analyses, large scale proteomic studies of lysine acetylation sites revealed a previously unappreciated number of acetylated proteins. The first of such studies described 37 acetylated proteins in the cytoplasmic fraction of Hela cells and 133 in mouse liver mitochondria5. In a more recent study using rat tissues, 15,474 modification sites on 4,541 proteins were identified6. This study revealed that the majority of lysine-acetylated proteins localize to the cytoplasm (30%) and nucleus (30%), with the remainder found in mitochondria (15%), plasma membrane (15%), endoplasmic reticulum or Golgi apparatus (5%), and extracellular region (5%)6. Here we will highlight examples of lysine acetylation regulating cellular processes. Lysine acetylation plays a crucial role in the regulation of cytoskeleton-associated proteins (e.g., actin, tubulin, small GTPases). Stress fibers, composed of beta- and gamma-actin, are important for cell shape and movement. In muscle cells, alpha-actin constitutes the microfilaments and together with myosin assures the traction forces required for muscle contraction and also for cytoplasmic streaming in non-muscle cells. All three actin isoforms can be acetylated5,7. Also, several regulatory proteins of the actin cytoskeleton are modified by acetylation. For example, several subunits of the Arp2/3 complex, important for actin nucleation, are acetylated7. Other studies have identified nine different lysine residues in cortactin which can be acetylated by HAT P300 or PCAF. Acetylation of those residues inhibits cortactin’s translocation to the cell periphery and reduces its binding capacity to actin, leading to diminished actin dynamics and altered cell motility8,9. Conversely, cortactin deacetylation, mediated by growth factor-stimulated Rac1 activation, leads to cortactin’s translocation from the cytosol to the cell periphery, where it interacts with and enhances the formation of F-actin10,11. Rac1 and other Rho family GTPases have a crucial role in regulating actin dynamics that underlie formation of lamellipodia, filopodia, and actin stress fibers. The activation of these G-proteins is tightly regulated by a variety of proteins, including GDIs (GDP dissociation inhibitors). Acetylation of RhoGDI alpha prevents its inhibitory effect on Rho family members, leading to enhanced stress fiber and filopodia formation5.

RhoA proteins are also inhibited by p120 catenin, a protein found in complexes with cadherin adhesion molecules. The p120 catenin is acetylated on at least three lysines which alters its subcellular localization and is important for its inhibition of RhoA5,12. Like actin and Rho GTPases, the cytoskeletal protein tubulin is also acetylated, with the Lys40 residue on alpha-tubulin identified as the acetylation site3. Studies have shown that tubulin acetylation contributes to microtubule stabilization and allows for more efficient bundling of acetylation-stabilized microtubules13,14. Microtubules are important for cellular transport, especially in neuronal cells with long extensions like axons and dendrites. Both kinesin and dynein-dependent microtubular transport are enhanced by tubulin acetylation15.
Besides these structural proteins, signaling proteins unrelated to histone modification and transcriptional regulation also have lysine acetylation sites. These proteins include Mdm2, Ku70, Stat3, Smad7, and Hsp90. Mdm2 is a RING finger E3 ubiquitin ligase that promotes ubiquitination and proteasomal degradation of the p53 tumor suppressor protein. Acetylation of Mdm2 impairs its ability to promote p53 ubiquitination and subsequent proteasomal degradation16. Interestingly, p53 itself can also be acetylated, leading to the destabilization of p53-Mdm2 interactions, which enables the p53-mediated stress response17. Ku70 acetylation promotes the dissociation of Bax from the Ku70/Bax complex, freeing Bax to localize to the mitochondria to execute its apoptotic function18.
In addition to influencing protein function, lysine acetylation also regulates the cellular localization of proteins, especially for nuclear import and export. Interestingly, for some proteins, acetylation favors localization to the cytoplasm19,20, whereas for others, acetylation will enhance the retention of proteins in the nucleus21,22. The mechanism by which acetylation regulates cellular localization can be either the alteration of an interaction with a binding partner leading to localization in a particular compartment (e.g., p53-Mdm2 interaction), or a modified interaction with nuclear import/export factors. For example, acetylation of Lys239 within the carboxyl-terminal nuclear localization signal (NLS) in adenovirus-transforming protein (E1A) disrupts its interaction with importin-alpha, thereby enhancing its localization in the cytoplasmic compartment23. Conversely, acetylation of hepatocyte nuclear factor-4 (HNF-4) at the NLS is important for its retention in the nucleus, thus preventing its export to the cytoplasm via the CRM-1 pathway24.
Conclusion
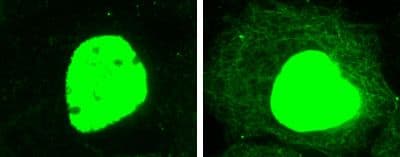
Lysine acetylation is well known to play a key role in regulating gene transcription and other DNA-dependent nuclear processes. Proteomics studies have identified many possible substrates of lysine acetylation and a large fraction of them resides in the cytoplasmic compartment, implicating their involvement in regulating important cellular pathways. However, the functions of many of them are still not clear. At Cytoskeleton, we have developed a mouse monoclonal pan-acetyl-lysine antibody to help researchers study lysine acetylation modification of their target proteins (Fig. 1). Applications of the antibody, including Western blotting, immunoprecipitation, immunofluorescence labeling, and ChIP, have been validated. Please contact us at [email protected] for more information.
Acetyl Lysine Antibody
This highly robust Anti-Acetyl Lysine Antibody recognizes a wide range of acetylated proteins in IP, WB, ChIP, and IF applications. This Anti-Acetyl Lysine Antibody has many advantages when compared to other commercially available antibodies on the market. See for yourself here.
Anti-Acetyl Lysine Antibody (Cat. # AAC01)
Acetyl Lysine Analysis Protocols
Optimized Acetyl Lysine Analysis Protocols Available Here
References
- Vidali G. et al. 1968. Chemical studies of histone acetylation. The distribution of epsilon-N-acetyllysine in calf thymus histones. J. Biol. Chem. 243, 6361-6366.
- Sterner R. et al. 1979. Studies of acetylation and deacetylation in high mobility group proteins. Identification of the sites of acetylation in HMG-1. J. Biol. Chem. 254, 11577-11583.
- L’Hernault S.W. and Rosenbaum J.L. 1985. Chalamydomonas alpha-tubulin is posttranslationally modified by acetylation on the epsilon-amino group of a lysine. Biochemistry. 24, 473-478.
- Gu W. and Roeder R.G. 1997. Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell. 90, 595-606.
- Kim S.C. et al. 2006. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell. 325, 607-618.
- Lundby A. et al. 2012. Proteomic analysis of lysine acetylation sites in rat tissues reveals organ specificity and subcellular patterns. Cell Rep. 2, 419-431.
- Choudhary C. et al. 2009. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science. 325, 834-840.
- Zhang X. et al. 2007. HDAC6 modulates cell motility by altering the acetylation level of cortactin. Mol. Cell. 27, 197-213.
- Zhang Y. et al. 2009. Deacetylation of cortactin by SIRT1 promotes cell migration. Oncogene. 28, 445-460.
- Weed S.A. et al. 1998. Translocation of cortactin to the cell periphery is mediated by the small GTPase Rac1. J. Cell Sci. 111, 2433-2443.
- Weed S.A. et al. 2000. Cortactin localization to sites of actin assembly in lamellipodia requires interactions with F-actin and the Arp2/3 complex. J. Cell Biol. 151, 29-40.
- Anastasiadis P.Z. et al. 2000. Inhibition of RhoA by p120 catenin. Nat. Cell Biol. 2, 637-644.
- Hubbert C. et al. 2002. HDAC6 is a microtubule-associated deacetylase. Nature. 417, 455-458.
- Matsuyama A. et al. 2002. In vivo destabilization of dynamic microtubules by HDAC6-mediated deacetylation. EMBO J. 21, 6820-6831.
- Dompierre J.P. et al. 2007. Histone deacetylase 6 inhibition compensates for the transport deficit in Huntington’s disease by increasing tubulin acetylation. J. Neurosci. 27, 3571-3583.
- Wang X. et al. 2004. Inhibition of p53 degradation by Mdm2 acetylation. FEBS Lett. 561, 195-201.
- Tang Y. et al. 2008. Acetylation is indispensable for p53 activation. Cell. 133, 612-626.
- Cohen H.Y. et al. 2004. Acetylation of the C terminus of Ku70 CBP and PCAF controls Bax-mediated apoptosis. Mol. Cell. 13, 627-638.
- Bari M.G. et al. 2006. C-Abl acetylation by histone acetyltransferases regulates its nuclear-cytoplasmic localization. EMBO Rep. 7, 727-733.
- Valacco M.P. et al. 2006. Cell growth-dependent subcellular localization of p8. J. Cell. Biochem. 97, 066-1079.
- Thevent L. et al. 2004. Regulation of human SRY subcellular distribution by its acetylation/deacetylation. EMBO J. 23, 3336-3345.
- Naryzhny S.N. and Lee H. The post-translational modifications of proliferating cell nuclear antigen: acetylation, not phosphorylation, plays an important role in the regulation of its function. J. Biol. Chem. 279, 20194-20199.
- Madison D.L. et al. 2002. Acetylation of the adenovirus-transforming protein E1A determines nuclear localization by disruption with importin-alpha. J. Biol. Chem. 277, 38755-38763.
- Soutglou E. et al. 2000. Acetylation regulates transcription factor activity at multiple levels. Mol. Cell. 5, 745-751.