FtsZ protein: S. aureus recombinant 6xHis-tagged
Cat. #FTZ02


Cat. #FTZ02



FtsZ is a highly conserved bacterial cytoskeletal protein that polymerizes to form a contractile ring at the future site of cell division, playing a pivotal role in cytokinesis. Its dynamic assembly and GTPase activity make it a key target for studying bacterial cell cycle regulation and developing novel antimicrobial agents.
Staphylococcus aureus FtsZ protein is produced in a bacterial expression system;
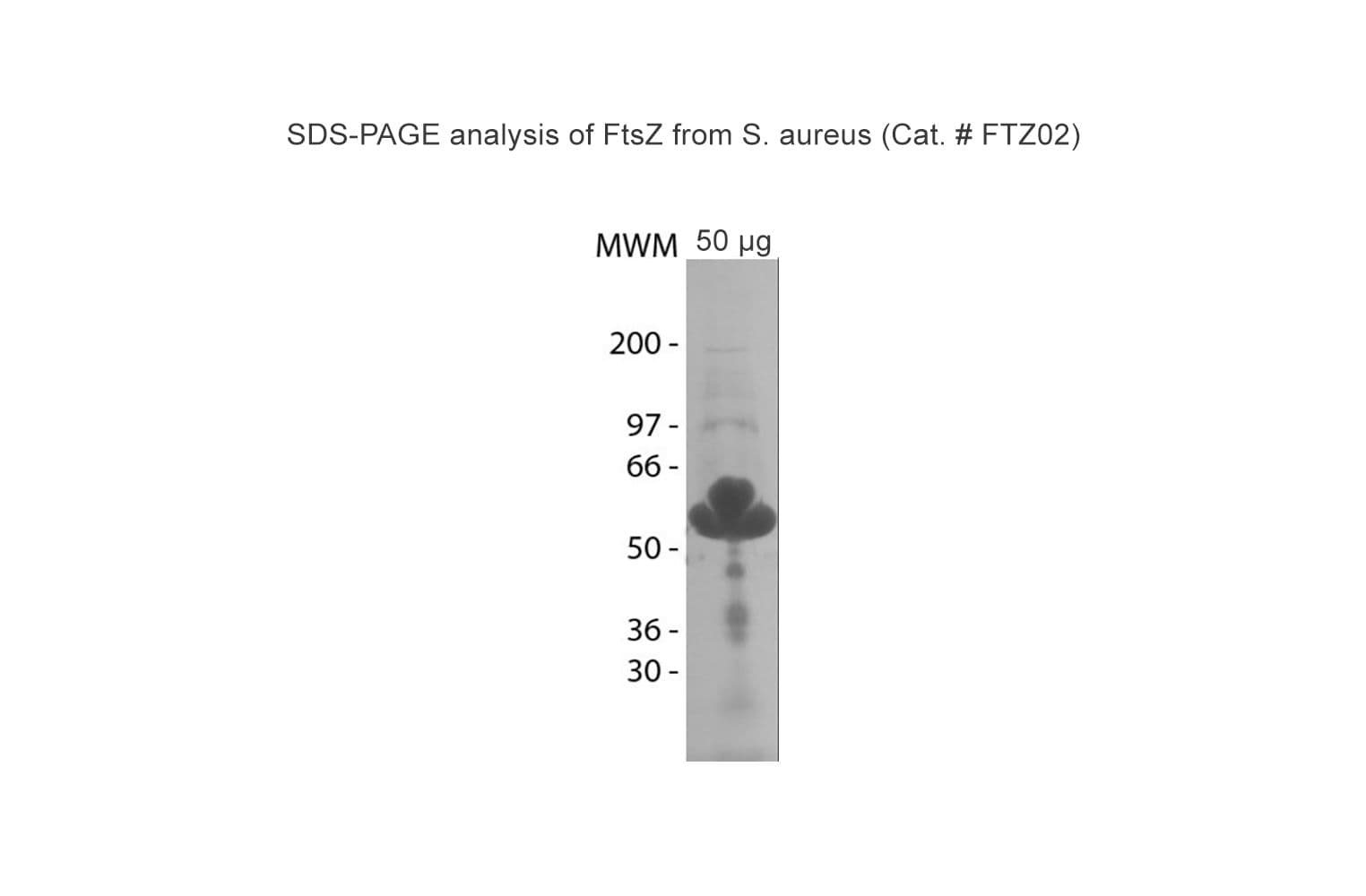
Protein purity is assessed by scanning densitometry of Coomassie Blue stained protein on a 4-20% polyacrylamide gel. Purity was determined to be ≥85% pure.
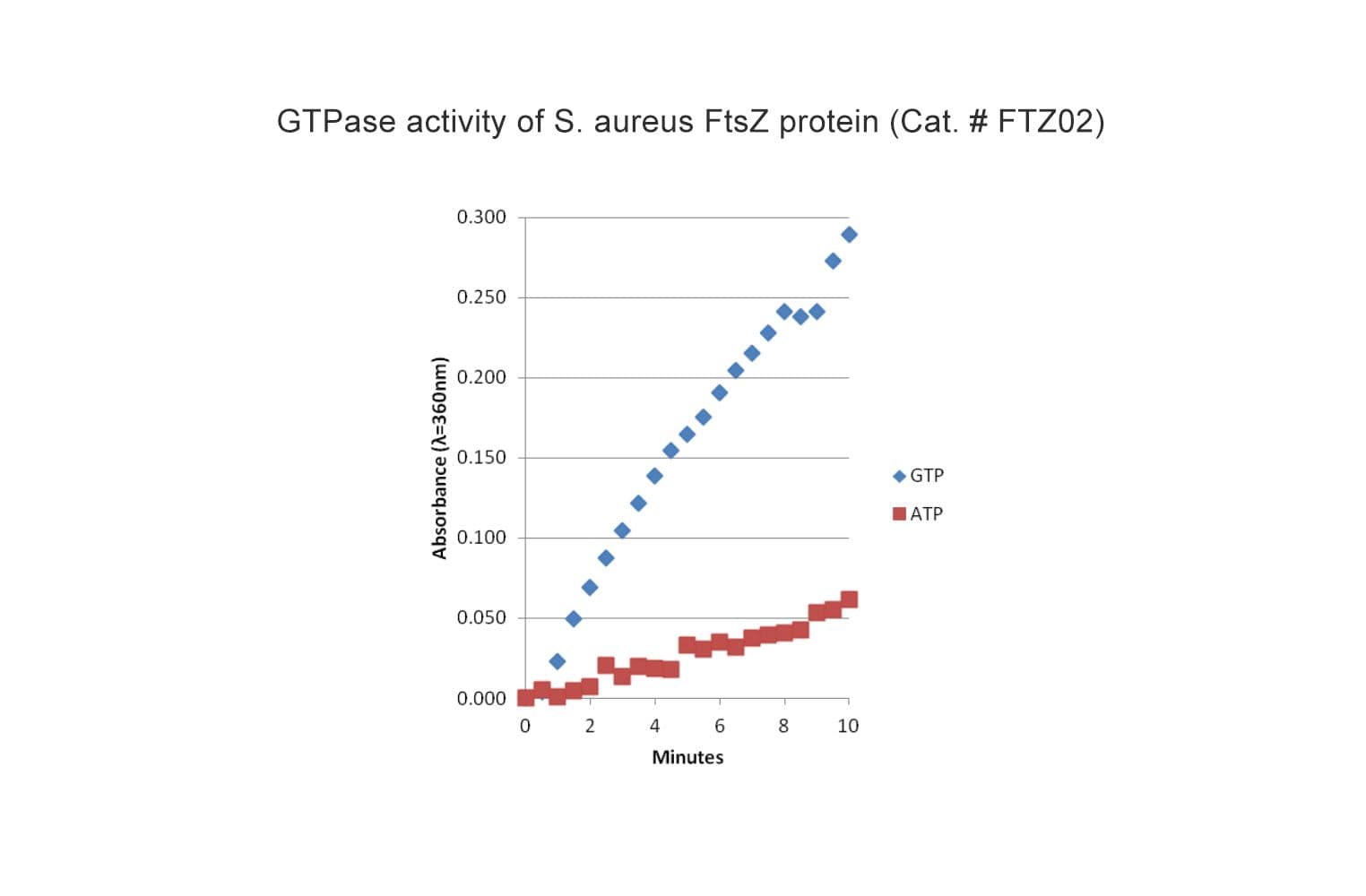
The biological activity of FTZ02 is determined by the ability of the protein to polymerize into protofilaments and sheets in vitro in the presence of Mg2+ and GTP. Under the experimental conditions (see datasheet) GTP-dependent S. aureus FtsZ polymerization achieves an OD360 between 0.30-0.40 after 10 minutes reaction time.