Phosphotyrosine (pTyr) post-translational modifications (PTMs) are crucial regulatory switches in cell signaling, controlling processes like growth, differentiation, and immune responses through reversible tyrosine phosphorylation. Dysregulation of these modifications is linked to diseases such as cancer, making them key therapeutic targets.
APY03-beads are an affinity reagent comprising a high-affinity/specificity pan-phosphotyrosine antibody APY03 covalently linked to agarose beads.
Key characteristics
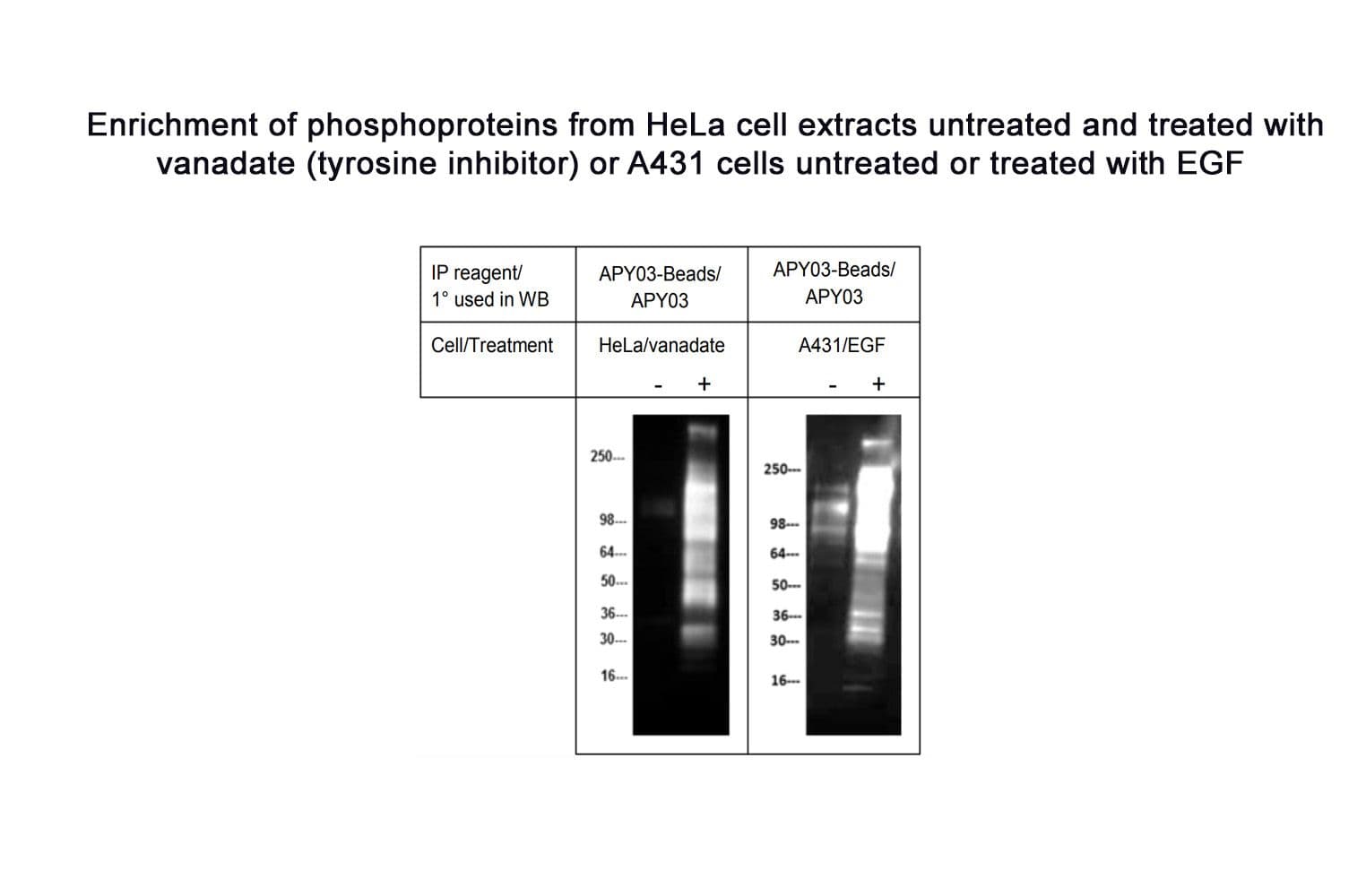
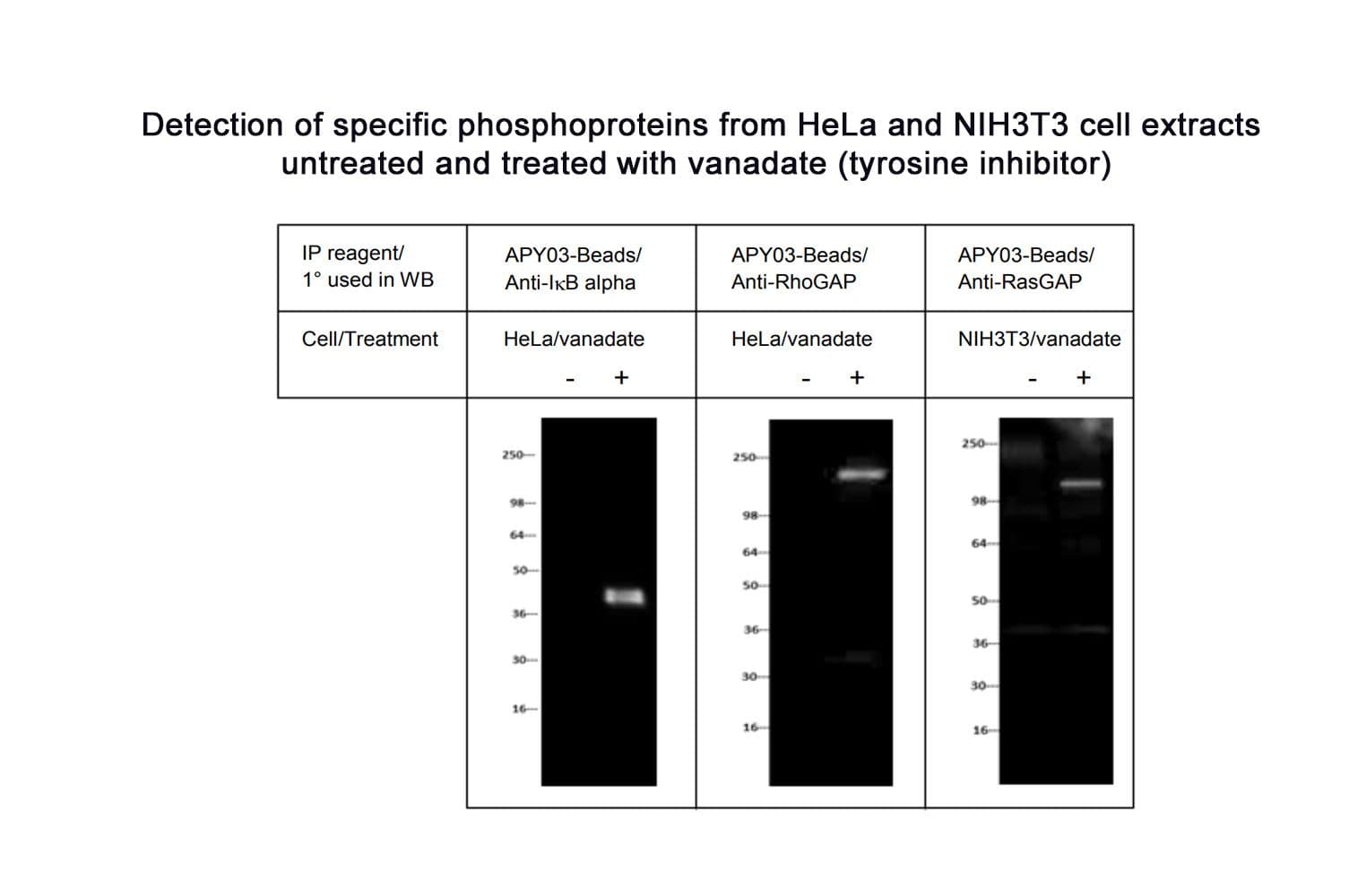
The biological activity of APY03-beads is demonstrated by the ability of the reagent to enrich a broad range of tyrosine phosphorylated proteins in HeLa cells treated with the protein tyrosine phosphatase inhibitor, pervanadate. The pTyr signal is enhanced ≥10-15 fold over untreated lysates (30 µl beads: 1mg lysate).
Cat. #APY03-Beads

© 2026 Cytoskeleton, Inc All Rights Reserved.