Arf1 G-LISA Activation Assay Kit (Colorimetric Based) - 96 assays
G-LISA Arf1 Activation Assay Biochem Kit (Colorimetric Based)
Product Uses Include
- Arf1 signaling pathway studies
- Arf1 activation assays with primary cells
- Studies of Arf1 activators and inactivators
- Arf1 activation assays with limited material
- High throughput screens for Arf1 activation
Introduction
With the new Arf1 G-LISA® kit you can now measure Arf1 activation from cell and tissue samples in less than 3 h. G-LISA® requires only 1-5% of the material needed for a conventional pull-down assay. You will also be able to handle large sample numbers and generate quantitative results. For a more detailed introduction on G-LISA® assays and a listing of other available G-LISA® kits, see our main G-LISA page.
The Arf1 G-LISA® kit contains an Arf-GTP-binding protein linked to the wells of a 96 well plate. Active, GTP-bound Arf1 in cell/tissue lysates will bind to the wells while inactive GDP-bound Arf1 is removed during washing steps. The bound active Arf1 is detected with a Arf1 specific antibody. The degree of Arf1 activation is determined by comparing readings from activated cell lysates versus non-activated cell lysates.
Kit contents
The kit contains sufficient reagents to perform 96 Arf1 activation assays. Since the Arf-GTP affinity wells are supplied as strips and the strips can be broken into smaller pieces, each kit can be used for anywhere from one to multiple assays. The following components are included in the kit:
- 96 well Arf1-GTP binding plate (divisible into 12 strips of 8 wells each)
- Anti-Arf1 antibody
- HRP-labeled secondary antibody
- Arf1 control protein (constitutively active Arf1)
- Cell Lysis buffer
- Wash buffer
- Binding buffer
- Antigen presenting buffer
- Antibody dilution buffer
- HRP detection reagent A
- HRP detection reagent B
- HRP stop solution
- Precision Red™ advanced protein assay reagent (Cat. # ADV02)
- Protease inhibitor cocktail (Cat. # PIC02)
- Manual with detailed protocols and extensive troubleshooting guide
Equipment needed
- 96-well plate spectrophotometer capable of reading 490 nm wavelength
- Multichannel or multidispensing pipettor
- Orbital microplate shaker capable of at least 200 rpm shaking (400 rpm is optimal)
Example results
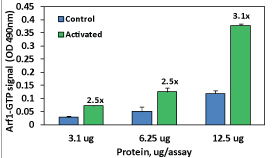
Figure 1. Arf1 activation measured by G-LISA. Lysates for Arf1 G-LISA were prepared from MDCK cells that were grown in serum-containing media and then either serum-starved for 1 hr (Activated) or remained in serum (Control). 12.5, 6.25, and 3.1 µg of cell lysates were subjected to the G-LISA assay. Absorbance was read at 490 nm. Data are background subtracted.
For product Datasheets and MSDSs please click on the PDF links below.
 G-LISA Activation Assay Technical Guide download here
G-LISA Activation Assay Technical Guide download here
Coming soon! If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com




