F-actin Visualization Biochem Kit (fluorescence format)
Product Uses Include
- Cellular colocalization with actin binding proteins by immunofluorescence.
- Detecting changes in actin morphology upon bacterial or viral infections.
- Detecting changes in actin morphology upon activation of small G-protein signal transduction pathways (see Fig. 1)
- Focal adhesion marker
- Marker for serum starvation of tissue culture cells (see Fig. 1)
Introduction
Conventional actins have a relative molecular mass of approximately 43 kDa. Monomeric actin (G-actin) can self-assemble (polymerize) into microfilaments (F-actin), the fundamental unit of the actin cytoskeleton. Actins are highly conserved within the eukaryotic kingdom and exist in higher eukaryotes as multigene families. Isoforms show distinct cellular and sub-cellular expression and localization. It has been demonstrated that different isoforms have subtly different biochemical properties in vitro which supports functional diversity within isotypes in vivo (1).
The actin cytoskeleton is a highly dynamic structure, a property under the tight regulation of more than 150 actin binding proteins (ABPs) (2, 3). It is involved in a large number of cellular processes, including muscle contraction, lamellopodia extrusion, cell locomotion, cytokinesis, intracellular transport and cytoplasmic streaming (1). The morphology of the actin cytoskeleton changes rapidly in response to a wide variety of internal and external stimuli. For example, figure 1 shows that calpeptin stimulation of serum starved 3T3 cells results in a rapid accumulation of actin stress fibers. This reponse is due to the activation of the small GTPase RhoA (4). As a further example, many pathogenic bacteria and viruses harness the host actin cytoskeleton for their intracellular spread, resulting in characteristic actin comet tails (5).
Fluorescent phalloidins selectively stain filamentous actin at nanomolar concentrations (6). They are the reagent of choice for F-actin staining of fixed cells for several reasons:
- Bind in a stoichiometric ratio of one phalloidin to one actin monomer
- Do not bind to monomeric G-actins (unlike many actin antibodies) which results in cleaner filament staining
- Binding properties do not change with actins from a wide variety of species
- Binding properties do not change between different actin isotypes
- Non-specific staining is negligible
Kit contents
This kit contains enough reagents for staining 300 coverslips (12 mm circular). The following components are included:
- Rhodamine Phalloidin (Cat. # PHDR1)
- Fixation Buffer
- Permeabilization Buffer
- Wash buffer
- Mounting Medium
- Dark Box
- Manual with detailed protocols and extensive troubleshooting guide.
Equipment needed
- Fluorescence microscope equipped with filters for rhodamine detection
- Microscope slides
Example results
Figure 1 below shows 3T3 cells under serum starvation conditions (A) and calpeptin stimulated conditions (B) stained with the F-actin Visualization Biochem Kit. It can be seen that most actin stress fibers disappear under serum starvation conditions, due to the de-activation of the Rho signaling pathway, while calpeptin stimulation of the cells results in a rapid accumulation of actin stress fibers due to the activation of Rho (4).
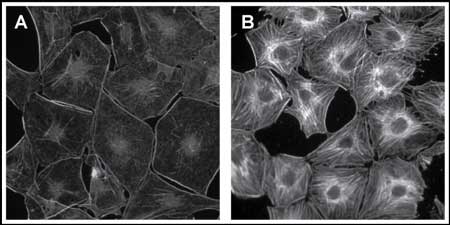
Figure 1. Phalloidin stained serum starved and calpeptin treated cells. Swiss 3T3 cells were grown in DMEM plus 10% calf bovine serum to 60% confluency, mediium was changed to 0.5% calf bovine serum for 24 h then to 0% calf bovine serum for an additional 24 h. After this serum starvation procedure, one dish was treated with calpeptin (0.1 mg/ml final) for 30 min while the other dish treated with carrier only (DMSO). After incubation, the coverslips were removed from the dishes and stained according to the BK005 manual. A: serum starved cells, B: calpeptin treated cells).
References
- Sheterline, P., Clayton, J., & Sparrow, J.C (Eds.). (1998) Protein Profile: Actin. 4th edition. Oxford University Press.
- dos Remedios, C.G. et al. (2003) Actin binding proteins: Regulation of cytoskeletal microfilaments. Physiol. Rev. 83: 433-473.
- Maciver S.K. The encyclopaedia of actin-binding proteins (and drugs)
- Ridley, A. J. and Hall, A. (1992) The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell 70: 389-399 (1992).
- Merz, A. J. and Higgs, H. N. (2003) Listeria Motility: Biophysics Pushes Things Forward. Curr. Biol. 13: R302–R304.
- Wulf, E., et al. (1979) Fluorescent phallotoxin: a tool for the visualization of cellular actin. Proc. Natl. Acad. Sci. USA 76: 4498–4502.
For product Datasheets and MSDSs please click on the PDF links below. For additional information, click on the FAQs tab above or contact our Technical Support department at tservice@cytoskeleton.com
Question 1: Do I need to use the fixative included with the F-actin visualization Biochem Kit or can I substitute something else?
Answer 1: For successful phalloidin staining, F-actin must be fixed and maintained in its native conformation. Paraformaldehyde (or some derivative, usually at 4% concentration) is an excellent fixative for this purpose. Methanol is not a good choice for phalloidin staining because F-actin’s conformation is lost with that fixative.
Question 2: Which filter settings should I use to visualize rhodamine-phalloidin?
Answer 2: The phalloidin in the F-actin visualization biochem kit (Cat. # BK005) is labeled with rhodamine, so the fluorescence microscope will require an excitation filter of 535 nm and an emission filter of 585 nm.
Question 3: Can I make my own anti-fade media?
Answer 3: The mounting media provided in the kit contains antifade so it’s not needed for the kit protocol, however if another antifade is desired we suggest the ProLong™ and ProLong Gold™ anti-fade reagents from Life Technologies Inc. Another option is PVA-Dabco, available from Fluka Biochemica. All of these will work well with fluorescent phalloidin.
You can also make your own anti-fade reagent. Here are two different recipes.
Recipe 1
250 nM glucose oxidase
64 nM catalase
40 mM d-glucose in water
1% BME
These reagents can be combined in Tris or PBS buffer
Tip: Store all components, separately, at -20°C in aliquots of 10 μl at a 100X concentration. Catalase and glucose oxidase are dissolved in a Tris or PBS buffer and snap frozen in liquid nitrogen. When thawed and stored on ice, aliquots will maintain activity for several hours. Once mixed together use the solution within 1 h. For consistency, do not refreeze thawed aliquots.
Tip: Add the glucose oxidase last and just before actually using the imaging buffer. This will initiate the first part of the reaction, which depletes the oxygen from solution.
Tip: DTT can be substituted for BME. Use a final DTT concentration of 10mM. Note, however, BME and DTT have both been demonstrated to have negative effects on some dyes. This will need to be determined empirically.
Recipe 2
PVA-Dabco
For 25 ml of 2.5% PVA-DABCO (Make 25 ml of PVA-DABCO in 50 ml culture tube to allow ample room for mixing).
1. Using a 50 ml culture tube, weigh 6 gm of glycerol into the culture tube on the balance
2. Add 2.4 gm of PVA
3. Mix well by repeatedly inverting the tube until the PVA is coated with glycerol
4. Add 6 ml of distilled water and repeat mixing until relatively uniform
5. Mix overnight using a rotator at room temperature
6. Add 12 ml of 0.2 M Tris-HCl at pH8-8.5
7. Heat to 50°C in a water bath with mixing (approximately 30 min)
8. Add 0.625 gm of DABCO and mix well
9. Centrifuge at 5000g for 15 min
10. Remove supernatant, aliquot (suggest 1 ml), and store at -20°C
The solution may be kept up to 6 months at -20°C and up to one week at 4°C before
becoming milky. Do not refreeze.
Stock materials
PVA-polyvinyl alcohol Sigma (P8136)
DABCO- 1,4 diazabicyclo [2.2.2]octane Sigma (D2522)
Tris-HCl Sigma (T3253)
Glycerol Fisher G33-1
To make 0.2N Tris HCl:
500 ml H2O
15.76g Tris HCl
add 7 pellets NaOH
adjust pH to 8.0-8.5 .
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com










