VIDEO
+3
Loading...
Cat. #BK129

Kit contents (96 assays: strip-wells allow single or multiple assays per run)
Equipment & materials required
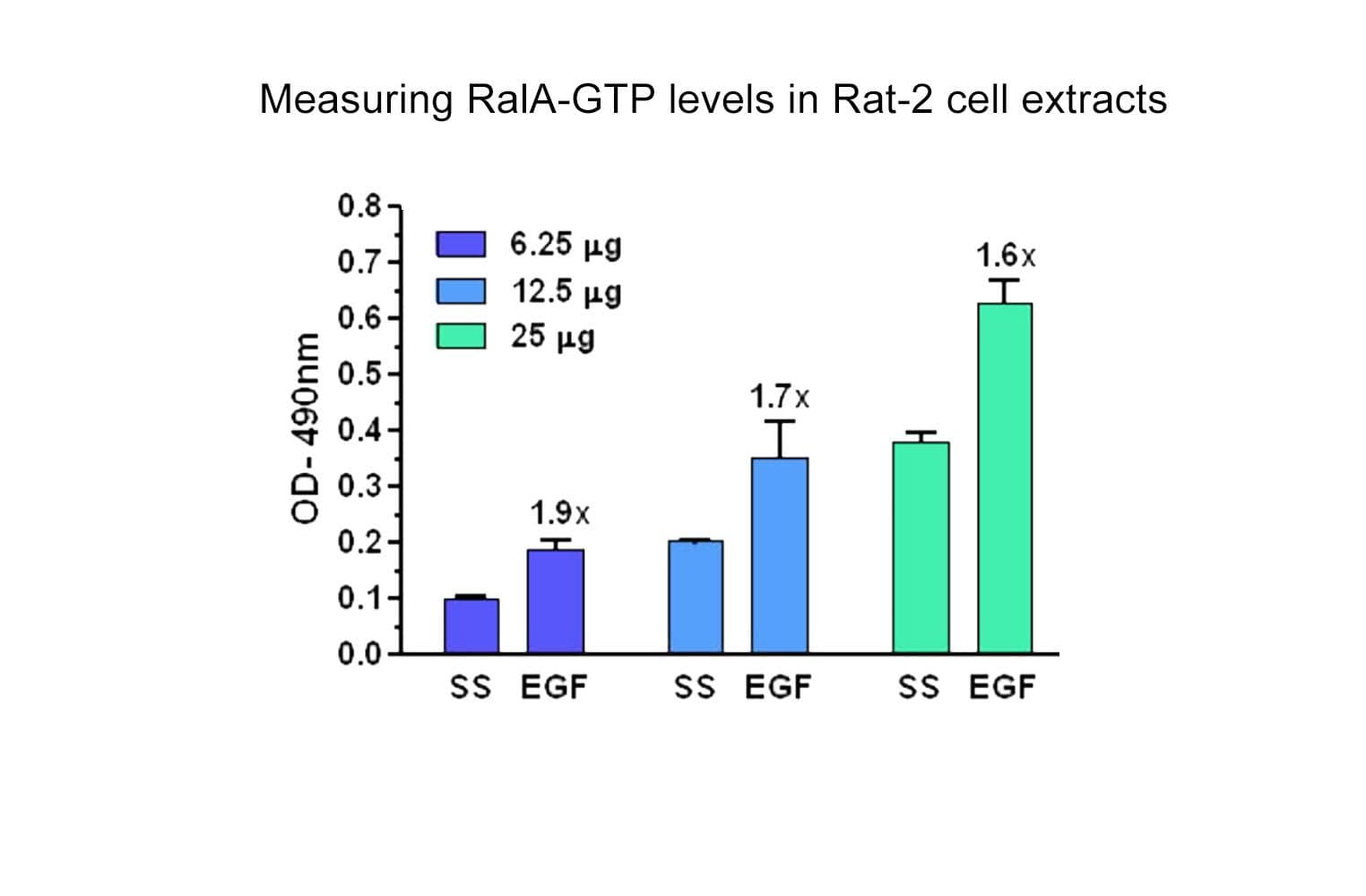
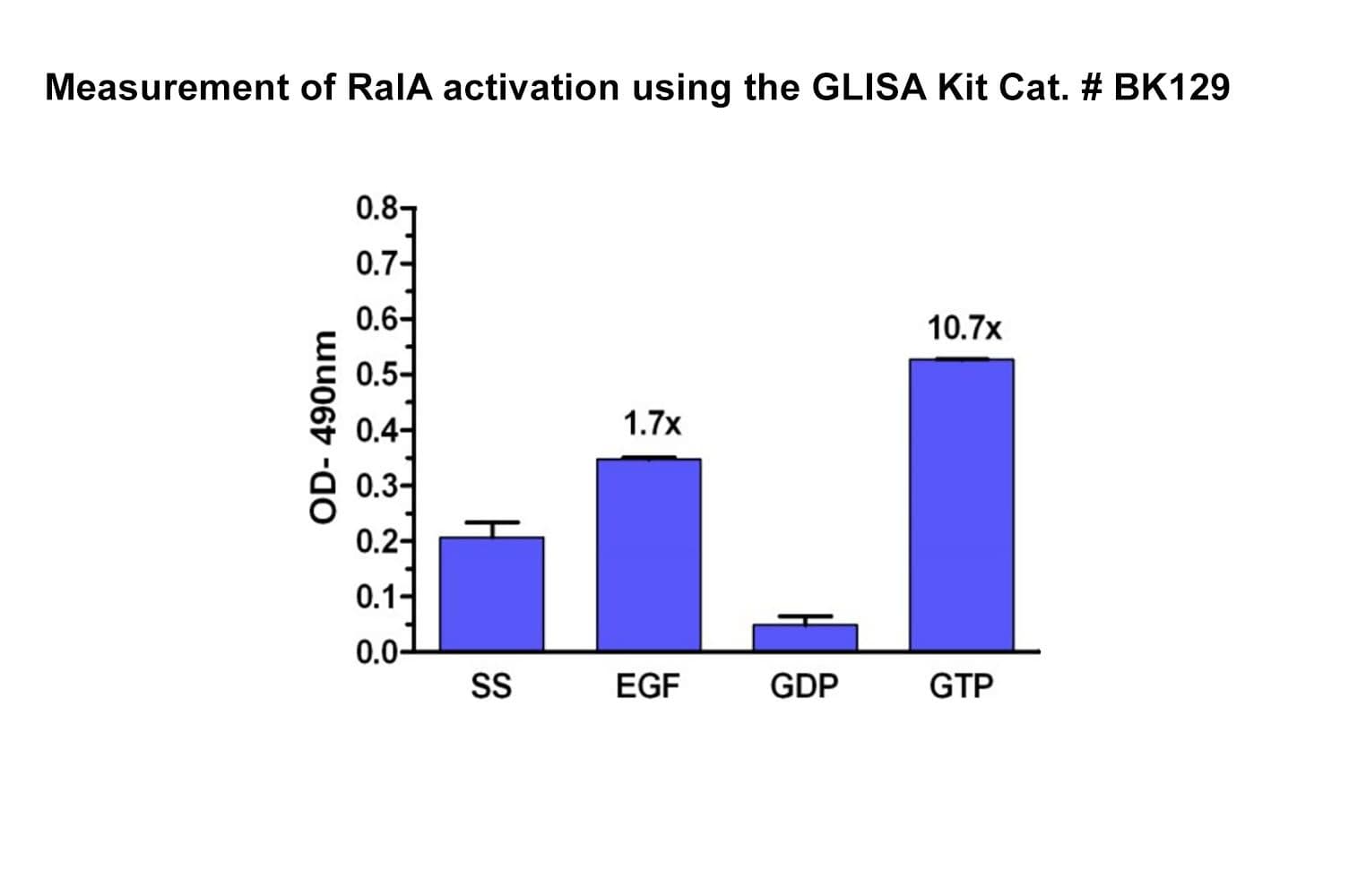
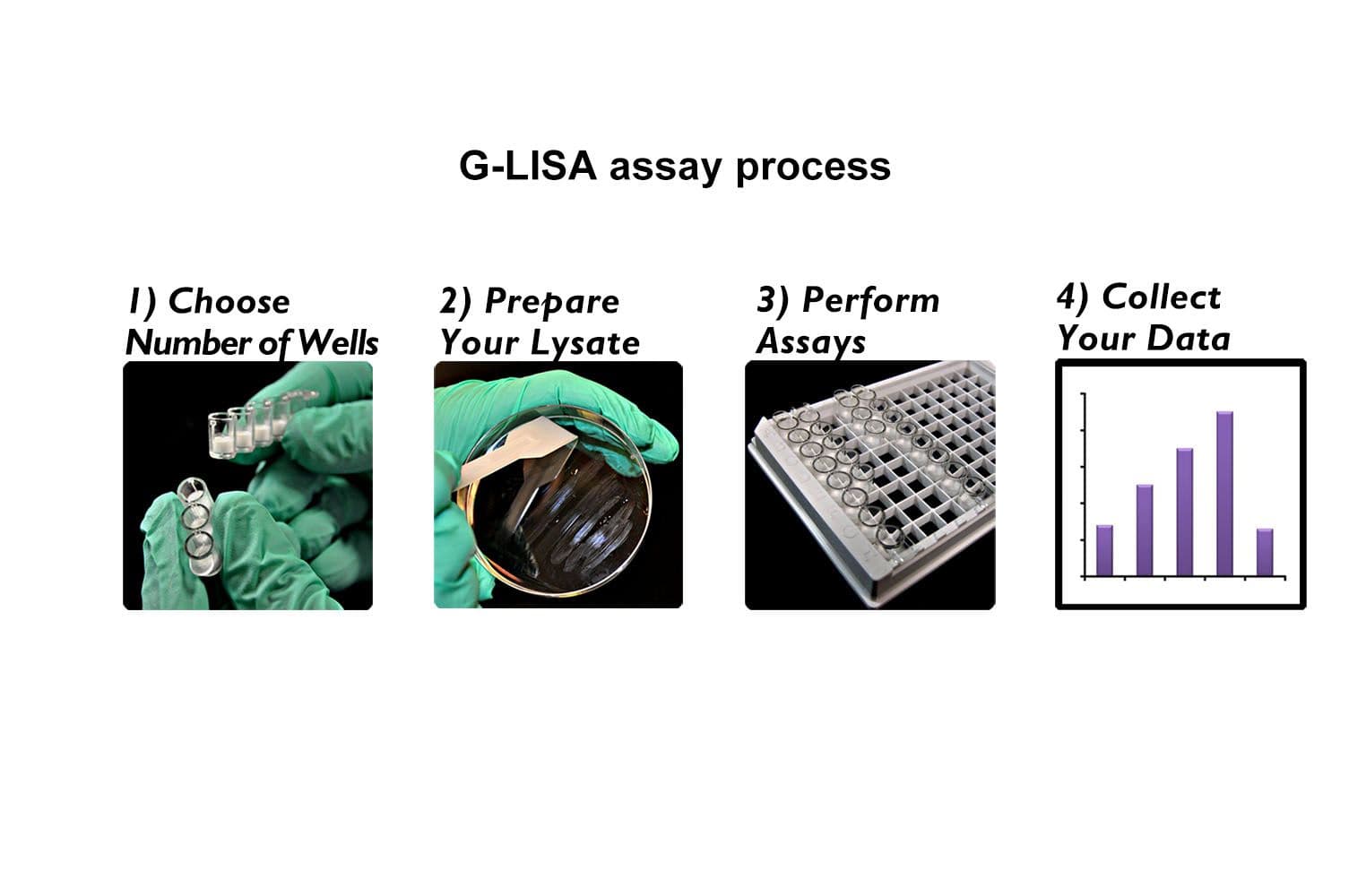
The Ral1 G-LISA® assay isolates active Ral (GTP-bound form) using the Ral-binding domain (RBD) of an effector protein coupled to a 96-well strip plate. The bound Ral is then detected and quantified using a colorimetric ELISA-based method and a RalA-specific antibody.
Key characteristics