VIDEO
+3
Loading...
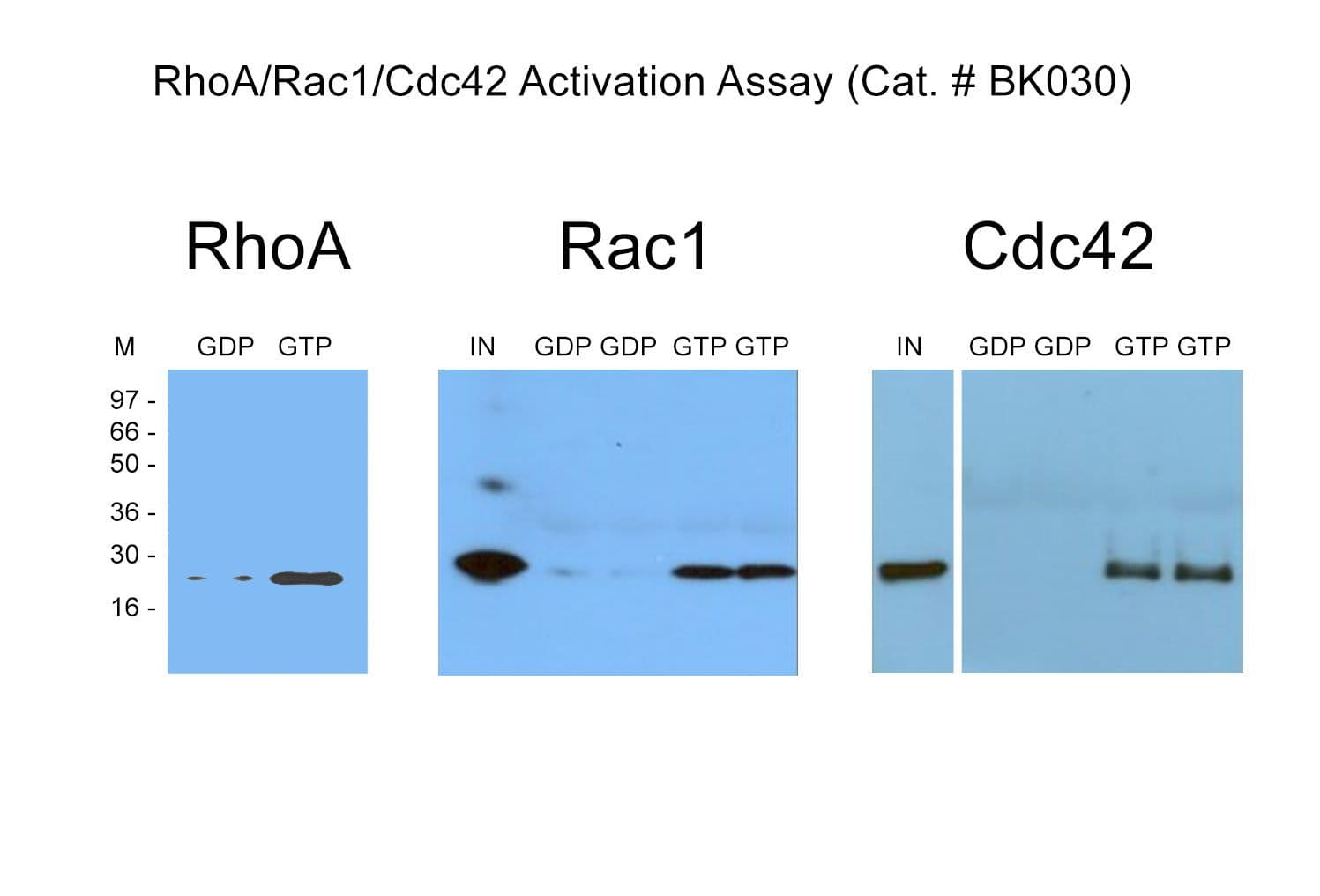
Cat. #BK030


Kit contents (3 x 10 assays)
Equipment & materials required
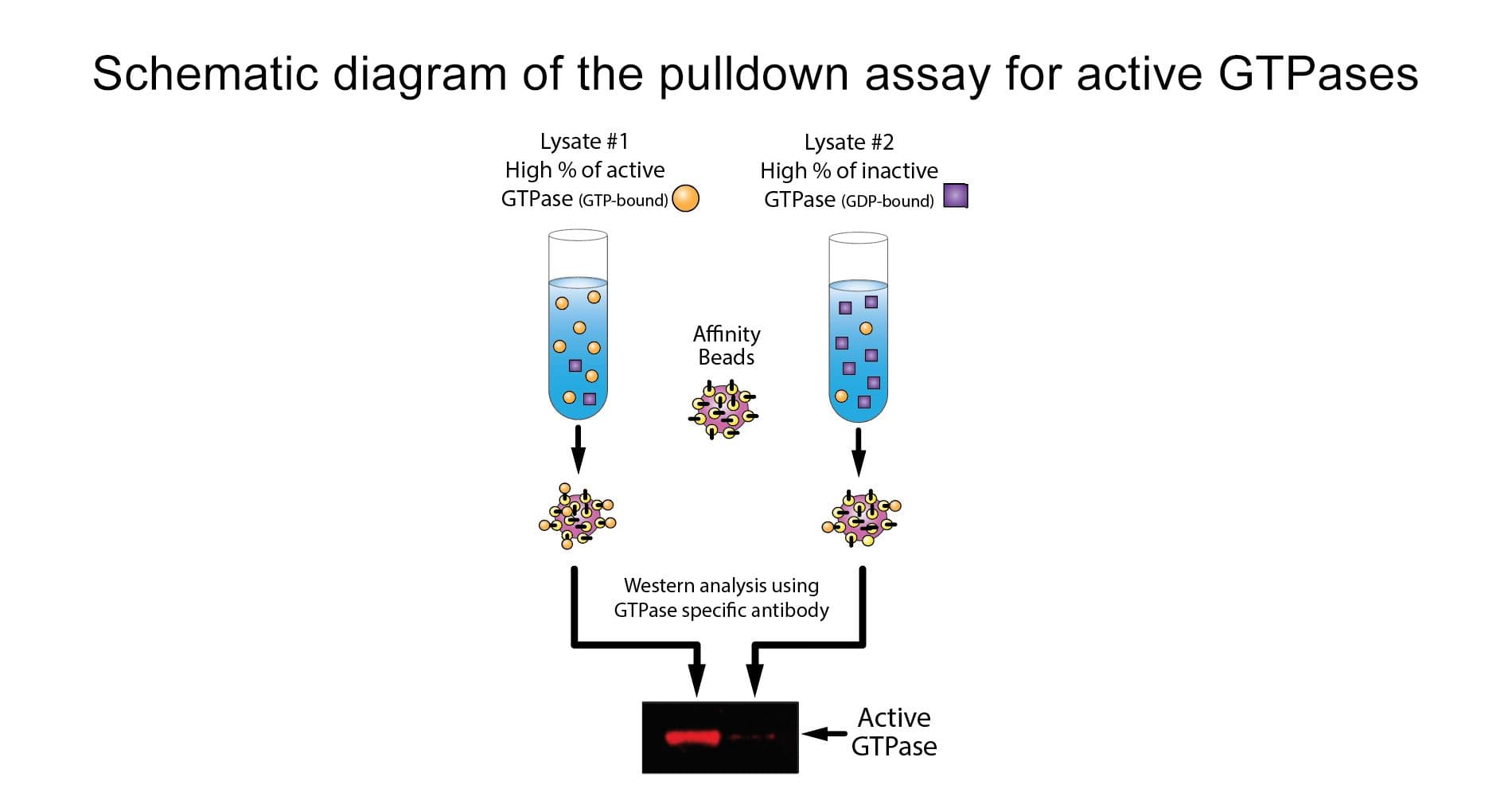
The BK030 kit is a combination sampler size of the Rac1, Cdc42 and RhoA pull-down kits. There is sufficient reagent to perform 10 pull-down assays for each GTPase.
Key characteristics