G-LISA® Small G-protein Activation Assays
G-LISA® – A revolutionary new way of doing Small G-protein Activation Assays!
There are multiple families of small GTPases that belong to the Ras GTPase superfamily. Among them are the Ras, Arf, and Rho families. The Ras and Arf families of small GTPases consist of many small G-proteins, including Ras and Ral isoforms in the former while Arf isoforms populate the latter. The Rho family of small GTPases consists of at least 20 members, the most extensively characterized of which are the RhoA, Rac1 and Cdc42 proteins. In common with other small GTPases, the Ras, Arf, and Rho proteins act as molecular switches that transmit cellular signals through downstream effector proteins by alternating between active GTP-bound and inactive GDP-bound states. The Ras, Arf, and Rho families mediate a wide range of cellular responses, including cytoskeletal reorganization, regulation of transcription, membrane trafficking and apoptosis. Therefore, understanding the mechanisms that regulate activation and inactivation of the GTPases is of great biological significance and is a subject of intense investigation.

The traditional way of measuring the level of activation of a Ras, Arf, and Rho protein has been to do so-called pull-down activation assays. However, these methods are time consuming, require large amount of sample, tend to not be very consistent and are not suitable for plate reader technology. To address these shortcomings, we have developed the G-LISA assay (Patent# 7,763,418 B2).
G-LISA Advantages (Click here)Activation Data (Click here)
Tech Notes(Click here)
Assays and Products(Click here)
The G-LISA Advantage
The G-LISA assay uses a 96-well plate coated with a binding domain of a Ras, Arf, or Rho-family effector protein. The active GTP-bound form of the Ras, Arf, or Rho-family protein, but not the inactive GDP-bound form, from a biological sample will bind to the plate, respectively. The bound active Ras, Arf, or Rho-family protein is then detected by incubation with a specific primary antibody followed by a secondary antibody conjugated to HRP. The signal is then developed with OPD or chemiluminescence reagents. These assays are simple, fast (<3 hours), require only small amounts of sample (5-50 µg cell protein) and yield quantitative and accurate data (see Table 1).
G-LISA assays are not only improving current experimental design, but serving as enabling technology, to facilitate experiments that were not possible with traditional pull-downs. For instance, cell cultures in 3D matrices grown for 10 or more days can now be analyzed for GTPase activation (1). The G-LISA assays also provide an accuracy and sensitivity of detection that allows analyses of GTPase activity in preparations previously off-limits with pull-down assays (2,3).
Recent studies have directly compared the G-LISA activation assays with pull-downs and the advantages are clear:
G-LISA assays are superior due to the ability to use small amounts of protein (1,4,5), their greater sensitivity (1,3,5) and quantitative measurements (2).

| Table 1: Comparison of traditional pull-down assay with G-LISA | ||
G-LISA | Traditional Pull-down | |
Cell material per assay* | 10-50 µg protein | 500-2000 µg protein |
Assay time | <3 h | 10-12 h |
Lysate clarification needed** | No | Yes |
Sample handling | Up to 96 samples (or more) | Up to 10 samples |
Quantitative data*** | Yes | Semi |
* Cytoskeleton recommends 10 ug at the lower range; however, the technical lower limit of detection with cells over-expressing a small G-protein is 1 ug and is also suitable. | ||
The G-LISA kits contain all the reagents needed for the activation assay. All you need are your cell or tissue samples, a platform shaker and a microtiter plate compatible luminometer or spectrophotometer. The kit provides enough material for 96 assays. The affinity wells can be separated from each other, so you can run anywhere from 2-96 assays per experiment and each kit can be used for numerous separate experiments.
The G-LISA kits are available in either luminometric or colorimetric detection versions. The assays are identical except for the final detection step. In general: the luminometric assays are more sensitive, while the colorimetric assays have a slightly lower level of variance. See Table 2 for comparison of the two versions of the kit.
| Table 2: Comparison of chemiluminesce and absorbance based G-LISA | ||
Luminometric | Colorimetric | |
Assay Time | <3 h | <3 h |
Cell material per assay | 1-25 µg protein | 2-50 µg protein |
Measurement parameters | Luminometer - high gain and 100 ms read per well | Spectrophotometer |
Detection limit* | 0.25 ng small G-protein | 0.50 ng small G-protein |
Linear range of detection* | 0.025-1.0 ng | 0.05-2.0 ng |
cv of 8 replicates | 16% | 12% |
*Determined by titrating the amount of recombinant Rho added to the wells | ||
1. Keely et al., 2007. Methods Enzymol. v 426, p 27.
2. Oliver et al., 2011. Br J Cancer. v 104, p 324.
3. Tanaka et al., 2010. Biochem Biophys Res Commun. v 399, p 677.
4. Scott et al., 2007. J Invest Dermatol. v 127, p 668.
5. Moniz et al., 2008. Cell Signal. v 20, p 1762.
Activation Data
| We have tested the G-LISA kits on several mammalian cell lines and with different well-known Rho activating stimuli (see list of tested cell types on our G-LISA FAQs page). All kits give results that are similar to those seen with pull-down assays. Luminometric and colorimetric assays give comparable results. Typical results are shown in Figures 2, 3 and 4. |

Figure 4. Time course of RhoA activation by 10 µg/ml LPA, 100 µg/ml calpeptin (Cat.# CN01) and 1 µg/ml CNF (Cat.# CN03) in Swiss 3T3 cells. Serum-starved Swiss 3T3 cells were stimulated with activator for the times shown in the graph and RhoA activation was analyzed with G-LISA kit BK121. LPA stimulation leads to a rapid and transient activation of RhoA, which peaks at 2-3 min and quickly declines to basal levels. Activation by calpeptin leads to a signal increase between 10-30 min which declines to basal after 60 min. Activation by CNF leads to signal increase after 60 min which is stable for up to 12 h.
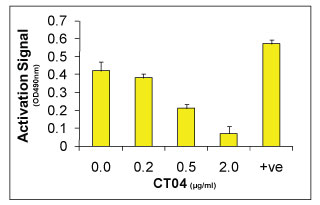
Figure 3. Rho activity measured in Swiss 3T3 cells treated with the Cell Permeable Rho Inhibitor (CT04) using the RhoA G-LISA Activation Assay (BK124). Serum starved Swiss 3T3 fibroblasts were untreated (no CT04) or treated with 0.20, 0.50 and 2.0 µg/ml of CT04 for 4 h in serum free medium at 37°C, then activated with 100 µg/ml calpeptin for 10 min. Cells were then lysed and RhoA activity was measured by the RhoA G-LISA Activation Assay (Cat.# BK124). Note: At 2.0 µg/ml CT04 for 4 h results in almost complete (90%) inhibition of RhoA activity.
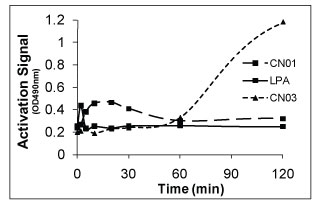
Figure 4. Time course of RhoA activation by 10 µg/ml LPA, 100 µg/ml calpeptin (Cat.# CN01) and 1 µg/ml CNF (Cat.# CN03) in Swiss 3T3 cells. Serum-starved Swiss 3T3 cells were stimulated with activator for the times shown in the graph and RhoA activation was analyzed with G-LISA kit BK121. LPA stimulation leads to a rapid and transient activation of RhoA, which peaks at 2-3 min and quickly declines to basal levels. Activation by calpeptin leads to a signal increase between 10-30 min which declines to basal after 60 min. Activation by CNF leads to signal increase after 60 min which is stable for up to 12 h.
Technical Tips
Click here for technical notes on the G-LISA assays
Click here for FAQs on the G-LISA assays
G-LISA Assays:
Arf1 G-LISA Activation Assay, colorimetric format (Cat.# BK132)
Arf6 G-LISA Activation Assay, colorimetric format (Cat.# BK133)
Rac1 G-LISA Activation Assay, luminescence format (Cat.# BK126)
Rac1 G-LISA Activation Assay, colorimetric format (Cat.# BK128)
Rac1,2,3 G-LISA Activation Assay, colorimetric format (Cat.# BK125)
RalA G-LISA Activation Assay, colorimetric format (Cat.# BK129)
Ras G-LISA Activation Assay, colorimetric format (Cat.# BK131)
RhoA G-LISA Activation Assay, colorimetric format (Cat.# BK124)
RhoA G-LISA Activation Assay, luminescence format (Cat.# BK121)
Cdc42 G-LISA Activation Assay, colorimetric format (Cat.# BK127)
Related Products:
Click here for products related to G-LISA activation assays
G-LISA® is a registered trademark of Cytoskeleton, Inc (CO). All rights reserved.
