Actin protein (biotin): skeletal muscle
Product Uses Include
- Microinjection into muscle cell followed by electron microscopy of streptavidin conjugated gold particles to determine the cellular localization of the modified actin.
- In vitro mot ility experiments using biotinylated seeds of F-actin bound to fluorescent beads for laser tweezer manipulation.
- Produce affinity columns using biotinylated F-actin or G-actin to extract and purify novel proteins.
- Purify proteins or develop bioassays using biotin F-actin or G-actin with streptavidin-magnetic beads.
Material
Rabbit skeletal muscle actin (Cat. # AKL99) has been modified to contain covalently linked biotin at random surface lysine residues. An activated ester of biotin is used to label the protein. The labeling stoichiometry has been determined to be approximately 1 biotin per actin monomer. Biotinylated actin has an approximate molecular weight of 43 kDa. AB07 is supplied as a lyophilized powder. The lyophilized protein when stored desiccated to < 10% humidity at 4°C is stable for 6 months. Th e protein should be reconstituted to 10 mg/ml with distilled water. The protein will then be in the following buffer: 5 mM Tris-HCl pH 8.0, 0.2 mM CaCl2, 0.2 mM ATP, 5% sucrose, and 1% dextran.
Purity
Protein purity is determined by scanning densitometry of Coomassie Blue stained protein on a 12% polyacrylamide gel. AB07 biotinylated actin is >99% pure. (see Figure 1.). No free biotin is apparent in the final product.
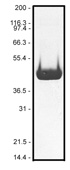
Figure 1. AB07 purity determination. A 10 µg sample of biotinylated actin (molecular weight approx. 43 kDa) was separated by electrophoresis in a 12% SDS-PAGE system. The protein was stained with Coomassie Blue. Protein quantitation was determined with the Precision Red Protein Assay Reagent (Cat.# ADV02).
Biological Activity
The biological activity of biotinylated actin is determined from its ability to efficiently polymerize into filaments in vitro and separate from unpolymerized components in a spin down assay. Stringent quality control ensures that 80-90% of the biotinylated actin can polymerized in this assay. This is comparable to the polymerization capacity of unmodified actin (Cat. # AKL99).
To determine the efficiency of biotin labeling, nanogram amounts of biotinylated actin are separated by electrophoresis and electroblotted onto a nitrocellulose membrane. The blot is then probed with streptavidin linked alkaline phosphatase. Quality control ensures that the biotin label on actin can be detected down to the 1 ng level of protein (see Figure 2). No free label is apparent in the final product.
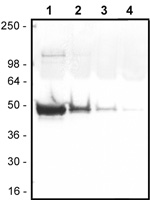
Figure 2. Detection of 1 ng of biotinylated actin. Serial dilutions of biotinylated actin were separated by electrophoresis on a 12% polyacrylamide gel and blotted onto Nitrocellulose. The membrane was then probed with streptavidin alkaline phosphatase (Sigma) and detected with the 1-Step NBT/BCIP reagent (Pierce). Lane 1: 1000 ng, Lane 2: 100 ng, Lane 3: 10 ng, and Lane 4: 1 ng of biotinylated actin.
For product Datasheets and MSDSs please click on the PDF links below. For additional information, click on the FAQs tab above or contact our Technical Support department at tservice@cytoskeleton.com
Question 1: Does biotinylated actin have the same polymerization dynamics as unlabeled actin?
Answer 1: The biological activity of biotinylated actin (Cat. # AB07) can be determined from its ability to efficiently polymerize into filaments in vitro and separate from unpolymerized components in a spin down assay. Stringent quality control ensures that 80-90% of the biotinylated actin can polymerize in this assay. This is comparable to the polymerization capacity of unmodified actin (Cat. # AKL99).
Question 2: Can the biotinylated actin be used in a pull-down format to capture novel actin binding proteins?
Answer 2: Yes, the biotinylated actin (Cat. # AB07) can be used to pull-down actin binding proteins with streptavidin beads. Below is a protocol:
1. Polymerize biotin actin at 0.4 mg/ml, using 20 mg AB07 plus 180 mg of unlabeled actin (Cat. # AKL99) for 1 h at RT, (alternatively for a monomer binding test, use AB07 diluted to 0.4 mg/ml in A-buffer for 1 h at RT, and add that to the beads 1 mg per 1 ml of beads)
2. Stabilize with 1 mM phalloidin (Sigma) added from 200 mM stock in methanol, diluted to 20 mM in F-actin buffer.
3. Mix with 200 ml packed volume of streptavidin beads washed with F-actin + phalloidin buffer. Incubate for 1 h at RT on a rotator 20 rpm, then wash again with 2 x 1 ml of F-actin + phalloidin buffer.
4. Use 20 mg AB07 per reaction = 20 ml of beads, plus 1 mg of cell extract protein. The assay uses the same binding conditions as in BK001, i.e., no more than 75 mM total ionic strength.
5.After 20 min incubation at RT, spin at 1000 rpm for 20 sec, pipette off supernatant, and save for a gel.
6. Wash the beads 2 x 1ml in binding buffer, then resuspend in 40 ml of 2x SDS loading buffer, heat to 95°C for 2 min and load 20 ml onto into the well, and load the supernatant sample next to it.
7. Three good controls are extract alone, streptavidin beads alone, and the monomer beads versus F-actin beads, all should be run with both pellet and supernatant samples.
If you have any questions concerning this product, please contact our Technical Service department at tservice@cytoskeleton.com