+3
Loading...
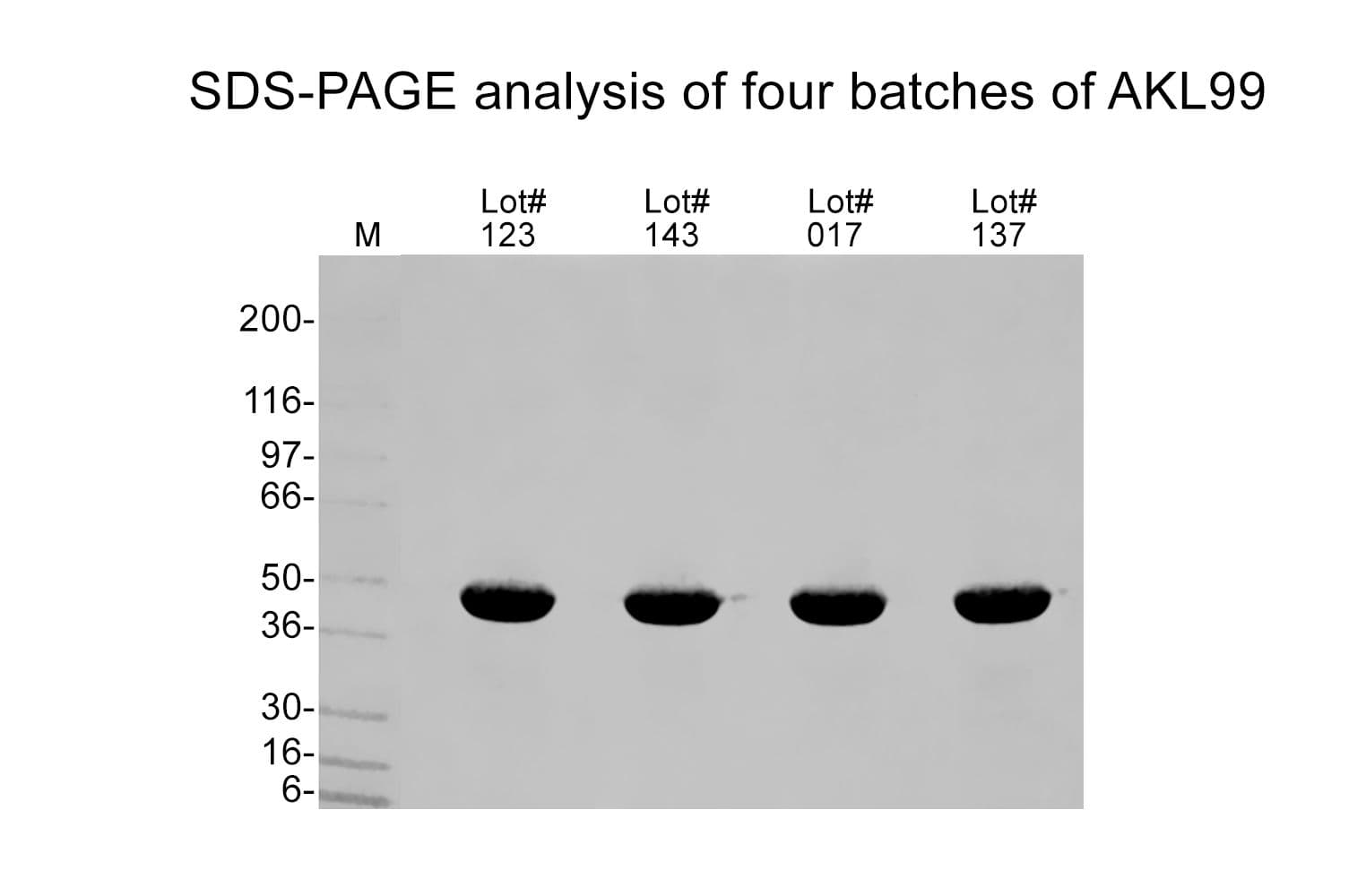
Alpha-skeletal actin protein has been purified from rabbit skeletal muscle and has an approximate molecular weight of 43 kDa. AKL99 is supplied as a white lyophilized powder.
Protein purity is assessed using scanning densitometry of Coomassie-stained SDS-PAGE gels. AKL99 is >99% pure alpha-skeletal muscle actin.
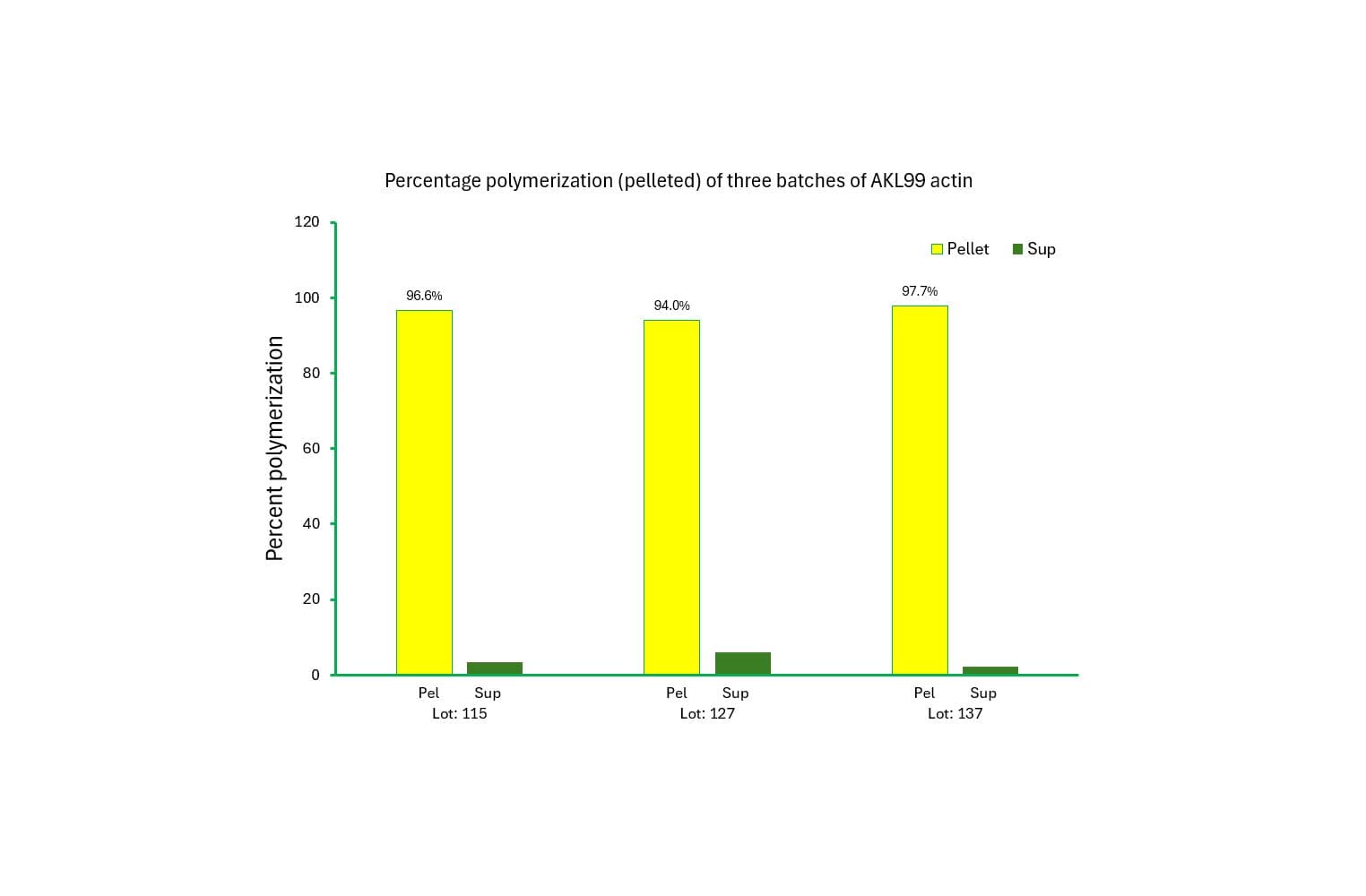
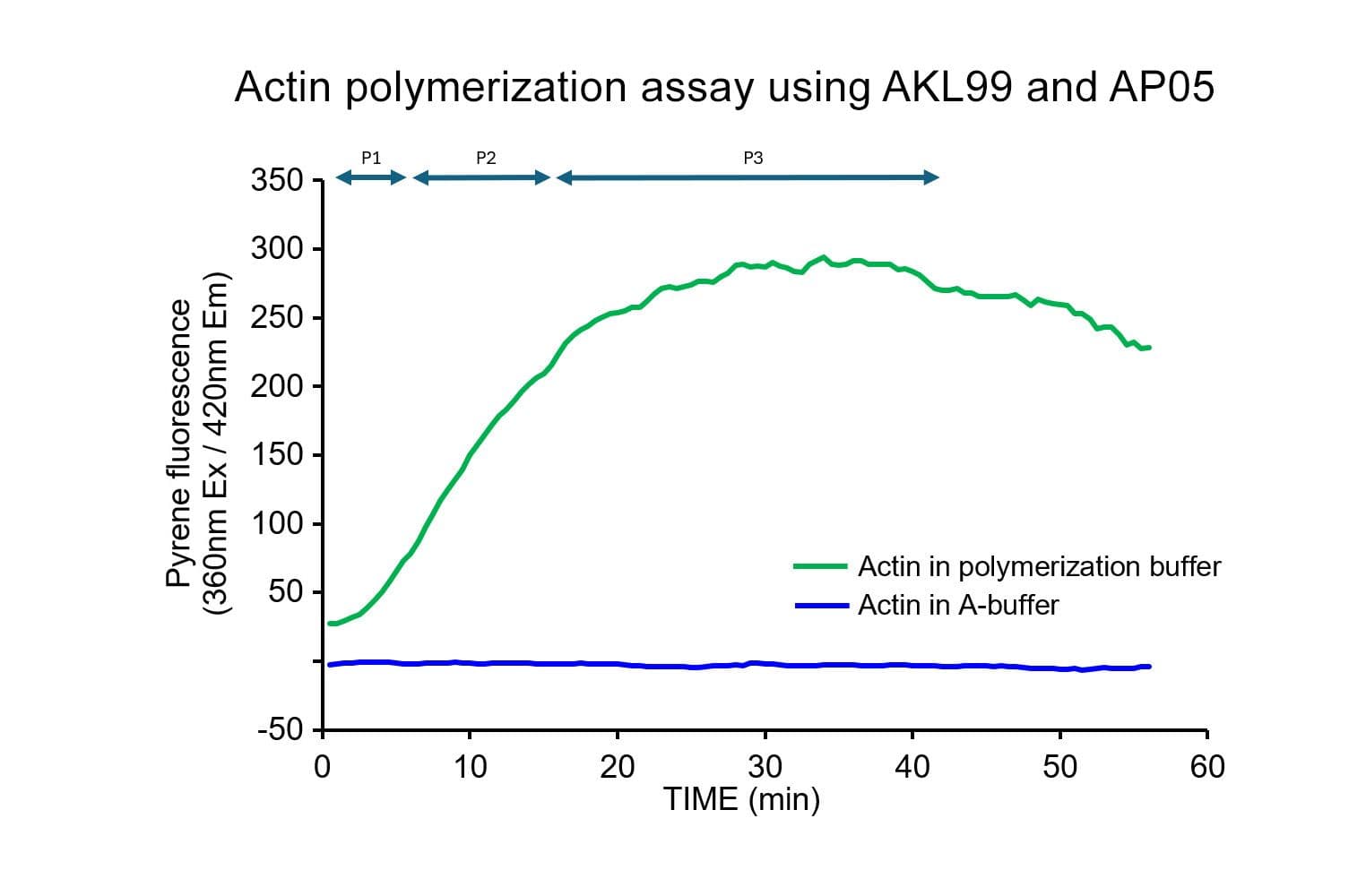
The biological activity of muscle actin is assessed by its ability to efficiently polymerize into filaments in vitro, which can be separated from unpolymerized components using a spin-down assay. Rigorous quality control ensures that >90% of the muscle actin polymerizes under these conditions.
Cat. #AKL99